Probe CUST_19442_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_19442_PI426222305 | JHI_St_60k_v1 | DMT400072782 | ACATATGTCTTGTTGTGCAACAAAAGTGCAGCATTTGACAAAGTTCAAAGGTAGTTTACC |
All Microarray Probes Designed to Gene DMG402028319
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_19380_PI426222305 | JHI_St_60k_v1 | DMT400072783 | ACATATGTCTTGTTGTGCAACAAAAGTGCAGCATTTGACAAAGTTCAAAGGTAGTTTACC |
| CUST_19442_PI426222305 | JHI_St_60k_v1 | DMT400072782 | ACATATGTCTTGTTGTGCAACAAAAGTGCAGCATTTGACAAAGTTCAAAGGTAGTTTACC |
Microarray Signals from CUST_19442_PI426222305
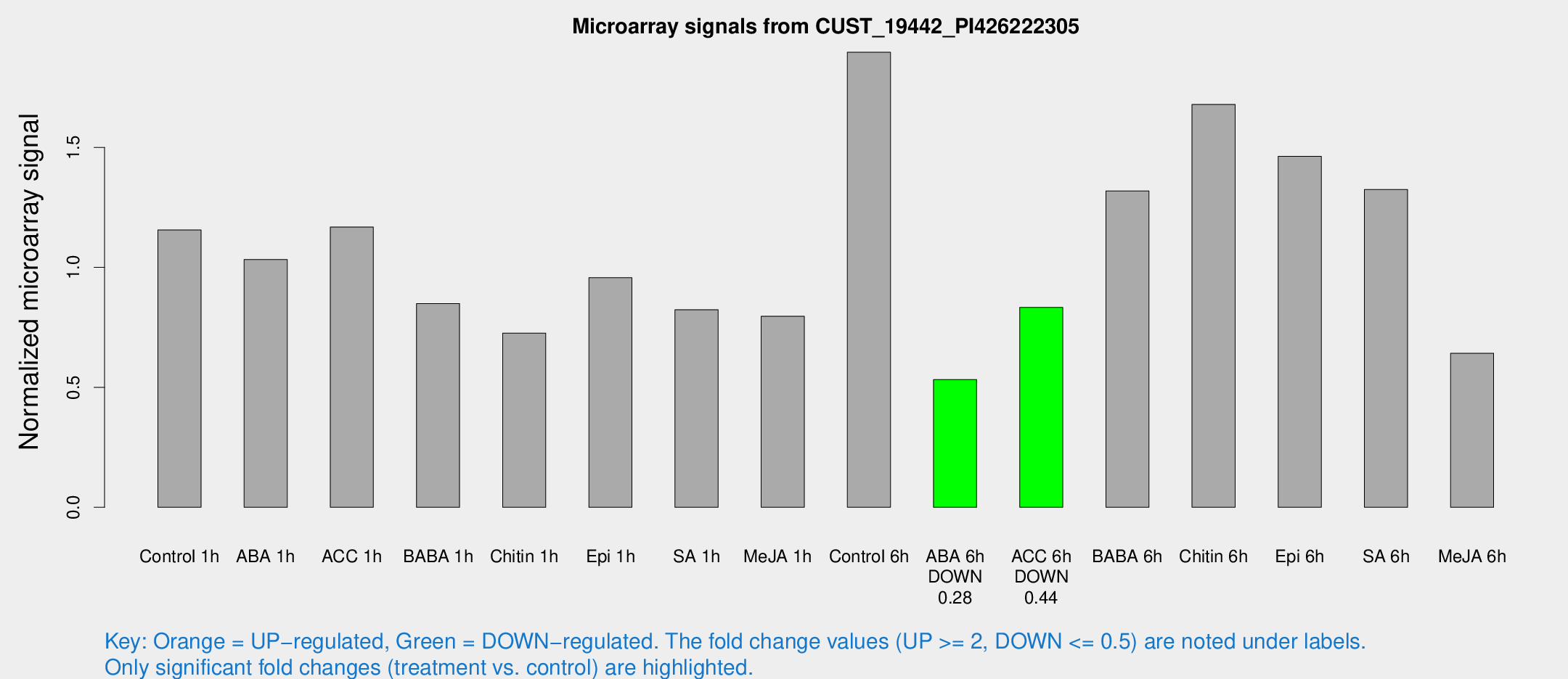
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 50.6633 | 9.72949 | 1.15589 | 0.154959 |
| ABA 1h | 39.7032 | 7.14994 | 1.03298 | 0.249037 |
| ACC 1h | 50.6118 | 4.27176 | 1.16828 | 0.0986789 |
| BABA 1h | 35.8423 | 7.27754 | 0.849034 | 0.151328 |
| Chitin 1h | 29.841 | 8.86527 | 0.725996 | 0.211833 |
| Epi 1h | 34.9518 | 3.51707 | 0.956768 | 0.0961552 |
| SA 1h | 35.8432 | 3.55635 | 0.823654 | 0.0823092 |
| Me-JA 1h | 27.5839 | 3.3526 | 0.796221 | 0.117645 |
| Control 6h | 81.9794 | 11.7544 | 1.89663 | 0.144109 |
| ABA 6h | 24.4672 | 3.91672 | 0.532415 | 0.0785376 |
| ACC 6h | 41.6503 | 7.86044 | 0.833058 | 0.0865349 |
| BABA 6h | 64.4702 | 12.5568 | 1.31862 | 0.250371 |
| Chitin 6h | 75.9844 | 6.82008 | 1.67927 | 0.122301 |
| Epi 6h | 70.3806 | 7.43813 | 1.46244 | 0.275853 |
| SA 6h | 57.5938 | 10.8483 | 1.32453 | 0.124567 |
| Me-JA 6h | 31.5315 | 10.932 | 0.641935 | 0.236545 |
Source Transcript PGSC0003DMT400072782 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G13820.1 | +2 | 3e-138 | 403 | 193/273 (71%) | alpha/beta-Hydrolases superfamily protein | chr1:4735622-4737617 FORWARD LENGTH=339 |
