Probe CUST_19380_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_19380_PI426222305 | JHI_St_60k_v1 | DMT400072783 | ACATATGTCTTGTTGTGCAACAAAAGTGCAGCATTTGACAAAGTTCAAAGGTAGTTTACC |
All Microarray Probes Designed to Gene DMG402028319
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_19380_PI426222305 | JHI_St_60k_v1 | DMT400072783 | ACATATGTCTTGTTGTGCAACAAAAGTGCAGCATTTGACAAAGTTCAAAGGTAGTTTACC |
| CUST_19442_PI426222305 | JHI_St_60k_v1 | DMT400072782 | ACATATGTCTTGTTGTGCAACAAAAGTGCAGCATTTGACAAAGTTCAAAGGTAGTTTACC |
Microarray Signals from CUST_19380_PI426222305
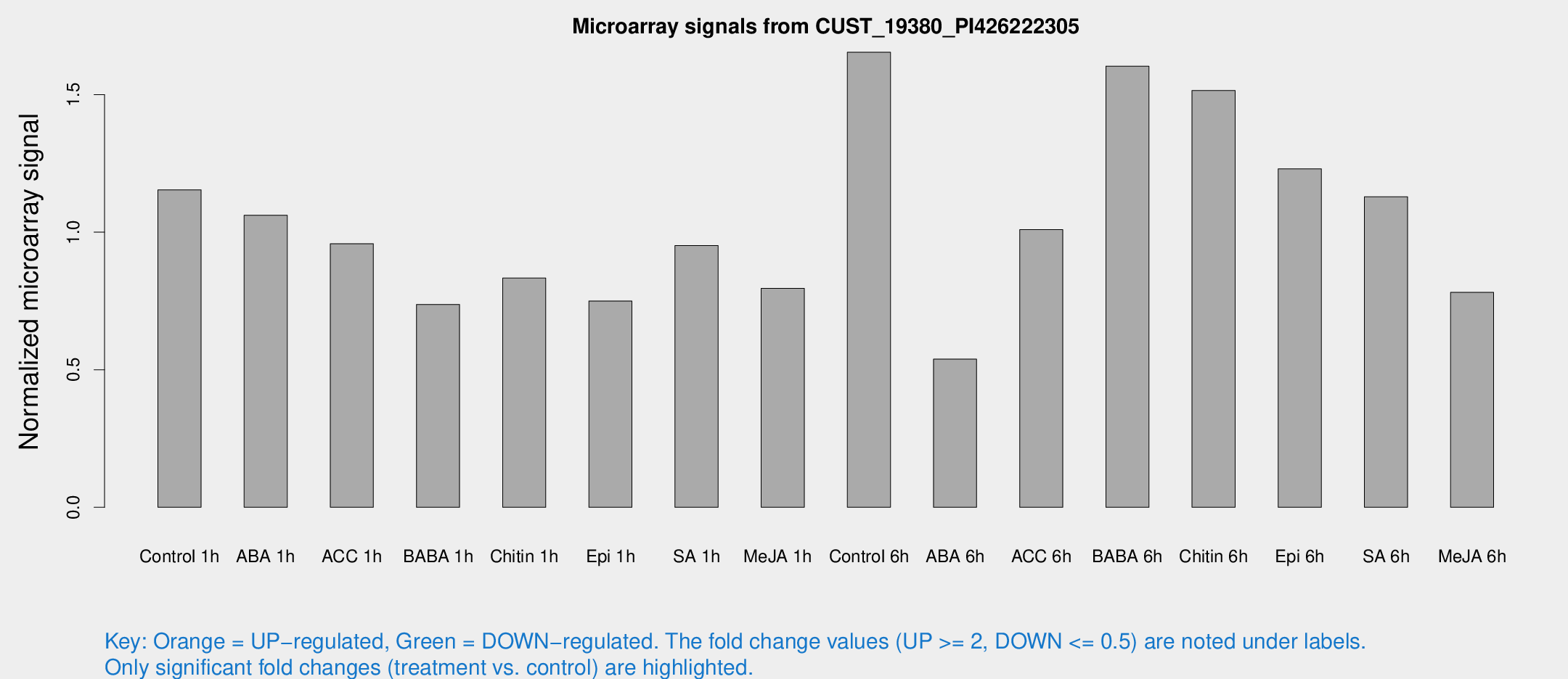
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 51.052 | 8.1384 | 1.15413 | 0.114095 |
| ABA 1h | 41.6591 | 6.66902 | 1.06169 | 0.192227 |
| ACC 1h | 43.2027 | 5.72719 | 0.957788 | 0.195507 |
| BABA 1h | 30.6647 | 3.92699 | 0.737276 | 0.109227 |
| Chitin 1h | 33.4953 | 5.92066 | 0.833454 | 0.118679 |
| Epi 1h | 28.7407 | 4.974 | 0.749786 | 0.133888 |
| SA 1h | 43.8574 | 8.95342 | 0.951623 | 0.262869 |
| Me-JA 1h | 28.4326 | 3.79618 | 0.795565 | 0.1338 |
| Control 6h | 73.2437 | 11.1274 | 1.65418 | 0.141084 |
| ABA 6h | 26.1818 | 5.76352 | 0.538688 | 0.178932 |
| ACC 6h | 50.8291 | 6.80611 | 1.00915 | 0.116294 |
| BABA 6h | 80.561 | 15.4105 | 1.60355 | 0.386693 |
| Chitin 6h | 69.9191 | 5.71423 | 1.51489 | 0.122829 |
| Epi 6h | 61.7378 | 10.4792 | 1.23024 | 0.333828 |
| SA 6h | 49.2597 | 7.88349 | 1.12879 | 0.169167 |
| Me-JA 6h | 37.8117 | 12.5784 | 0.78106 | 0.223116 |
Source Transcript PGSC0003DMT400072783 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G13820.1 | +3 | 3e-150 | 437 | 210/311 (68%) | alpha/beta-Hydrolases superfamily protein | chr1:4735622-4737617 FORWARD LENGTH=339 |
