Probe CUST_18834_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18834_PI426222305 | JHI_St_60k_v1 | DMT400001151 | ATGCCATTGAGGCTGTTCTTCTAATGATCGAAAATCCTGCCAGAGCCAATGGCCATATCT |
All Microarray Probes Designed to Gene DMG400000433
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18713_PI426222305 | JHI_St_60k_v1 | DMT400001150 | CTCCTACACCTACTCCTTTGTCCTAAATTTCATGCGCATGTTGTCGACACAATGTTTTTT |
| CUST_18834_PI426222305 | JHI_St_60k_v1 | DMT400001151 | ATGCCATTGAGGCTGTTCTTCTAATGATCGAAAATCCTGCCAGAGCCAATGGCCATATCT |
Microarray Signals from CUST_18834_PI426222305
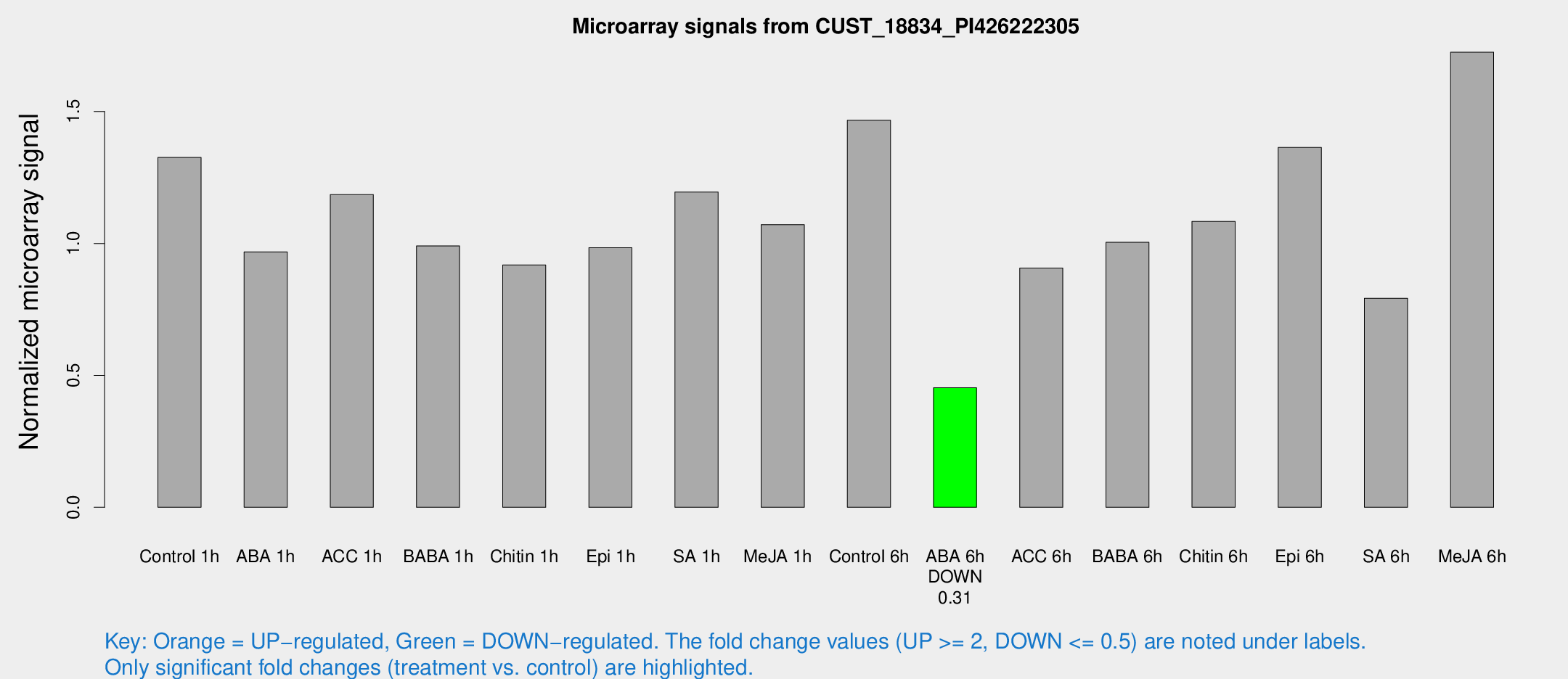
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 3883.07 | 559.836 | 1.32616 | 0.102917 |
| ABA 1h | 2485.95 | 258.083 | 0.968329 | 0.078697 |
| ACC 1h | 3711.67 | 815.523 | 1.18525 | 0.215882 |
| BABA 1h | 2781.22 | 319.31 | 0.9908 | 0.0572195 |
| Chitin 1h | 2378.14 | 137.72 | 0.918426 | 0.053042 |
| Epi 1h | 2471.76 | 247.958 | 0.983919 | 0.0853171 |
| SA 1h | 3568.1 | 410.451 | 1.19465 | 0.150375 |
| Me-JA 1h | 2517.36 | 145.713 | 1.07142 | 0.0763399 |
| Control 6h | 4694.73 | 1299.48 | 1.4673 | 0.393359 |
| ABA 6h | 1385.86 | 80.2539 | 0.453071 | 0.0345853 |
| ACC 6h | 3097.75 | 605.33 | 0.906755 | 0.0824844 |
| BABA 6h | 3258.36 | 321.638 | 1.00437 | 0.0947795 |
| Chitin 6h | 3338.14 | 289.48 | 1.08332 | 0.0721629 |
| Epi 6h | 4601.07 | 953.804 | 1.36408 | 0.323285 |
| SA 6h | 2365.54 | 483.672 | 0.792148 | 0.0933323 |
| Me-JA 6h | 5023.76 | 648.144 | 1.72496 | 0.14 |
Source Transcript PGSC0003DMT400001151 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G08200.1 | +1 | 0.0 | 723 | 351/385 (91%) | UDP-D-apiose/UDP-D-xylose synthase 2 | chr1:2574259-2576609 REVERSE LENGTH=389 |
