Probe CUST_18713_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18713_PI426222305 | JHI_St_60k_v1 | DMT400001150 | CTCCTACACCTACTCCTTTGTCCTAAATTTCATGCGCATGTTGTCGACACAATGTTTTTT |
All Microarray Probes Designed to Gene DMG400000433
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18713_PI426222305 | JHI_St_60k_v1 | DMT400001150 | CTCCTACACCTACTCCTTTGTCCTAAATTTCATGCGCATGTTGTCGACACAATGTTTTTT |
| CUST_18834_PI426222305 | JHI_St_60k_v1 | DMT400001151 | ATGCCATTGAGGCTGTTCTTCTAATGATCGAAAATCCTGCCAGAGCCAATGGCCATATCT |
Microarray Signals from CUST_18713_PI426222305
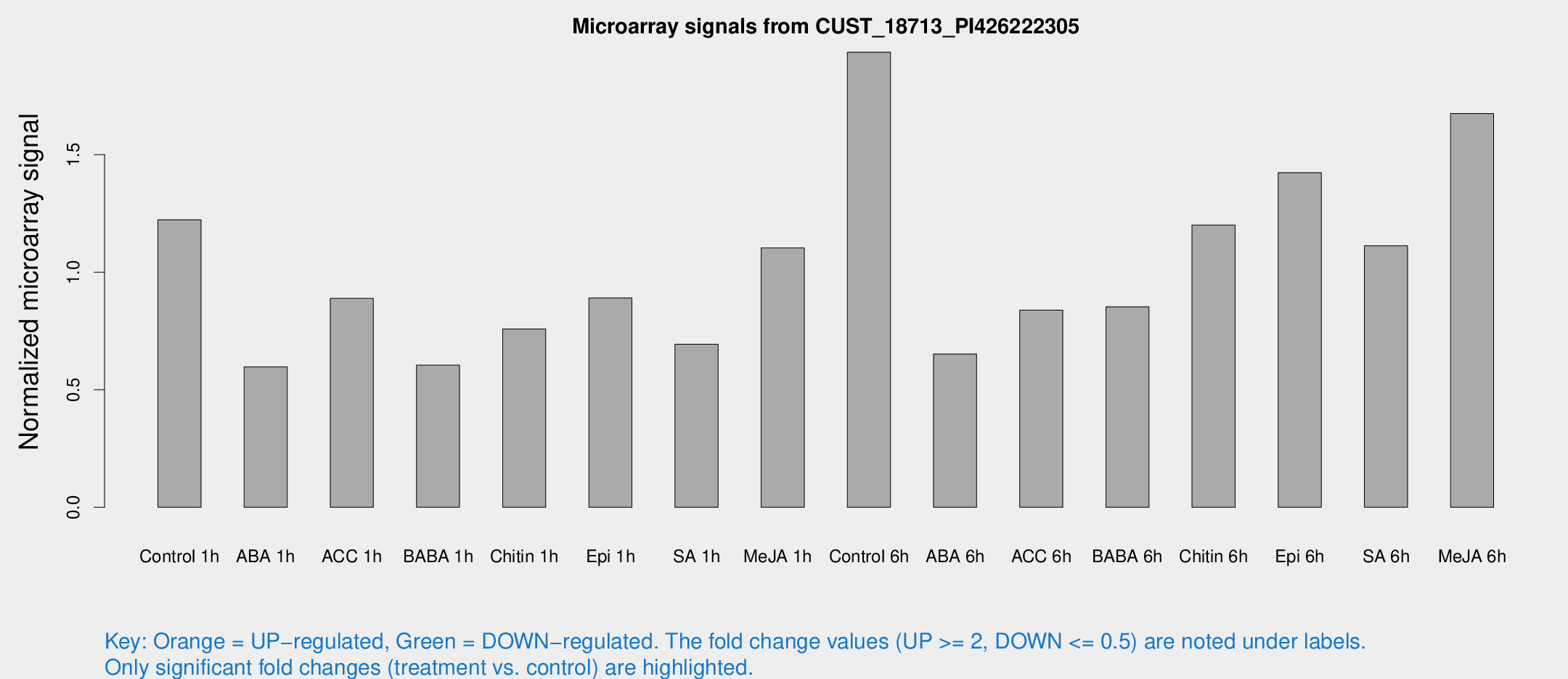
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 50.0735 | 4.81946 | 1.22263 | 0.117585 |
| ABA 1h | 23.2808 | 5.77558 | 0.59764 | 0.197195 |
| ACC 1h | 37.5783 | 4.79586 | 0.888584 | 0.11699 |
| BABA 1h | 26.8088 | 8.58527 | 0.604978 | 0.316035 |
| Chitin 1h | 32.5162 | 10.7847 | 0.758671 | 0.446819 |
| Epi 1h | 31.5792 | 3.85049 | 0.890497 | 0.109523 |
| SA 1h | 36.9807 | 13.9136 | 0.69341 | 0.561677 |
| Me-JA 1h | 37.9167 | 6.42607 | 1.103 | 0.162572 |
| Control 6h | 87.7054 | 24.4692 | 1.93568 | 0.488156 |
| ABA 6h | 31.0052 | 8.71205 | 0.651985 | 0.265259 |
| ACC 6h | 40.2226 | 5.72225 | 0.83865 | 0.134604 |
| BABA 6h | 40.8677 | 8.65522 | 0.852938 | 0.168085 |
| Chitin 6h | 52.8874 | 6.00949 | 1.20064 | 0.174219 |
| Epi 6h | 70.4497 | 18.4355 | 1.42353 | 0.475057 |
| SA 6h | 47.9876 | 12.8526 | 1.1124 | 0.454188 |
| Me-JA 6h | 75.97 | 23.4104 | 1.6746 | 0.449812 |
Source Transcript PGSC0003DMT400001150 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G08200.1 | +1 | 1e-179 | 518 | 251/272 (92%) | UDP-D-apiose/UDP-D-xylose synthase 2 | chr1:2574259-2576609 REVERSE LENGTH=389 |
