Probe CUST_18253_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18253_PI426222305 | JHI_St_60k_v1 | DMT400042190 | CCGTCGCCGGTGATGTTTCTCAGTCAGTGAACAATTGGATTTTCAAGCCAAAATTTGAGG |
All Microarray Probes Designed to Gene DMG401016370
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18253_PI426222305 | JHI_St_60k_v1 | DMT400042190 | CCGTCGCCGGTGATGTTTCTCAGTCAGTGAACAATTGGATTTTCAAGCCAAAATTTGAGG |
| CUST_18408_PI426222305 | JHI_St_60k_v1 | DMT400042189 | TCTCAGTCAGTGAACAATTGGATTTTCAAGCCAAAATTTGAGGGTAACAAAGAGATAAAG |
Microarray Signals from CUST_18253_PI426222305
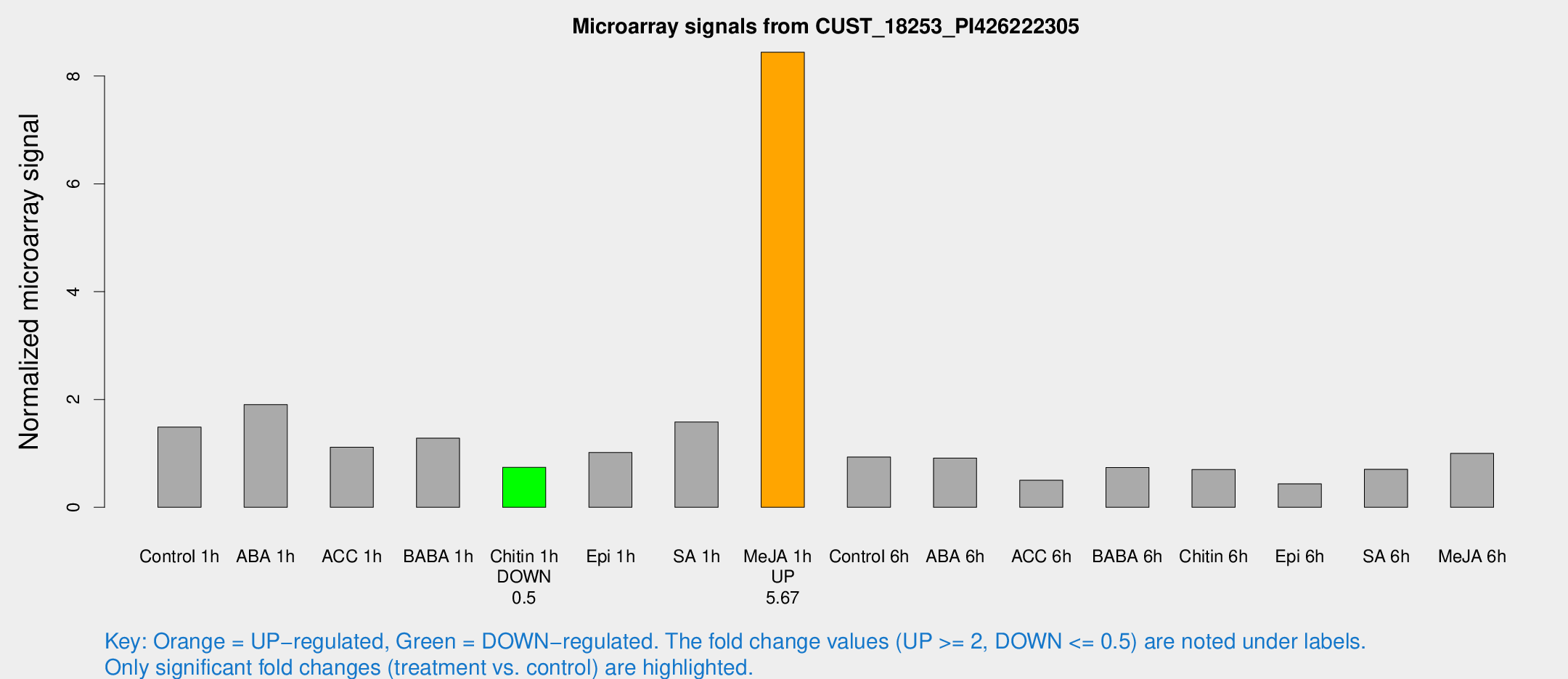
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 55.4721 | 9.25644 | 1.48855 | 0.16864 |
| ABA 1h | 66.854 | 20.8478 | 1.9067 | 0.447436 |
| ACC 1h | 45.4844 | 11.9277 | 1.1138 | 0.283922 |
| BABA 1h | 45.8904 | 7.0049 | 1.2813 | 0.118748 |
| Chitin 1h | 24.3308 | 3.23163 | 0.742644 | 0.0984056 |
| Epi 1h | 32.3218 | 3.44409 | 1.0168 | 0.111617 |
| SA 1h | 66.0641 | 20.2979 | 1.58315 | 0.697822 |
| Me-JA 1h | 251.222 | 15.209 | 8.44225 | 0.545589 |
| Control 6h | 38.7786 | 14.0905 | 0.932852 | 0.279291 |
| ABA 6h | 38.1693 | 9.75665 | 0.913286 | 0.232037 |
| ACC 6h | 21.3214 | 3.85534 | 0.501162 | 0.0892351 |
| BABA 6h | 33.4851 | 10.5607 | 0.738305 | 0.236451 |
| Chitin 6h | 29.9283 | 8.08816 | 0.701832 | 0.25938 |
| Epi 6h | 21.2323 | 7.08162 | 0.435447 | 0.285951 |
| SA 6h | 26.3224 | 5.20691 | 0.704536 | 0.169777 |
| Me-JA 6h | 41.5309 | 15.3468 | 0.999143 | 0.310084 |
Source Transcript PGSC0003DMT400042190 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G10740.1 | +1 | 0.0 | 640 | 292/376 (78%) | alpha/beta-Hydrolases superfamily protein | chr1:3568343-3570466 REVERSE LENGTH=473 |
