Probe CUST_18408_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18408_PI426222305 | JHI_St_60k_v1 | DMT400042189 | TCTCAGTCAGTGAACAATTGGATTTTCAAGCCAAAATTTGAGGGTAACAAAGAGATAAAG |
All Microarray Probes Designed to Gene DMG401016370
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18253_PI426222305 | JHI_St_60k_v1 | DMT400042190 | CCGTCGCCGGTGATGTTTCTCAGTCAGTGAACAATTGGATTTTCAAGCCAAAATTTGAGG |
| CUST_18408_PI426222305 | JHI_St_60k_v1 | DMT400042189 | TCTCAGTCAGTGAACAATTGGATTTTCAAGCCAAAATTTGAGGGTAACAAAGAGATAAAG |
Microarray Signals from CUST_18408_PI426222305
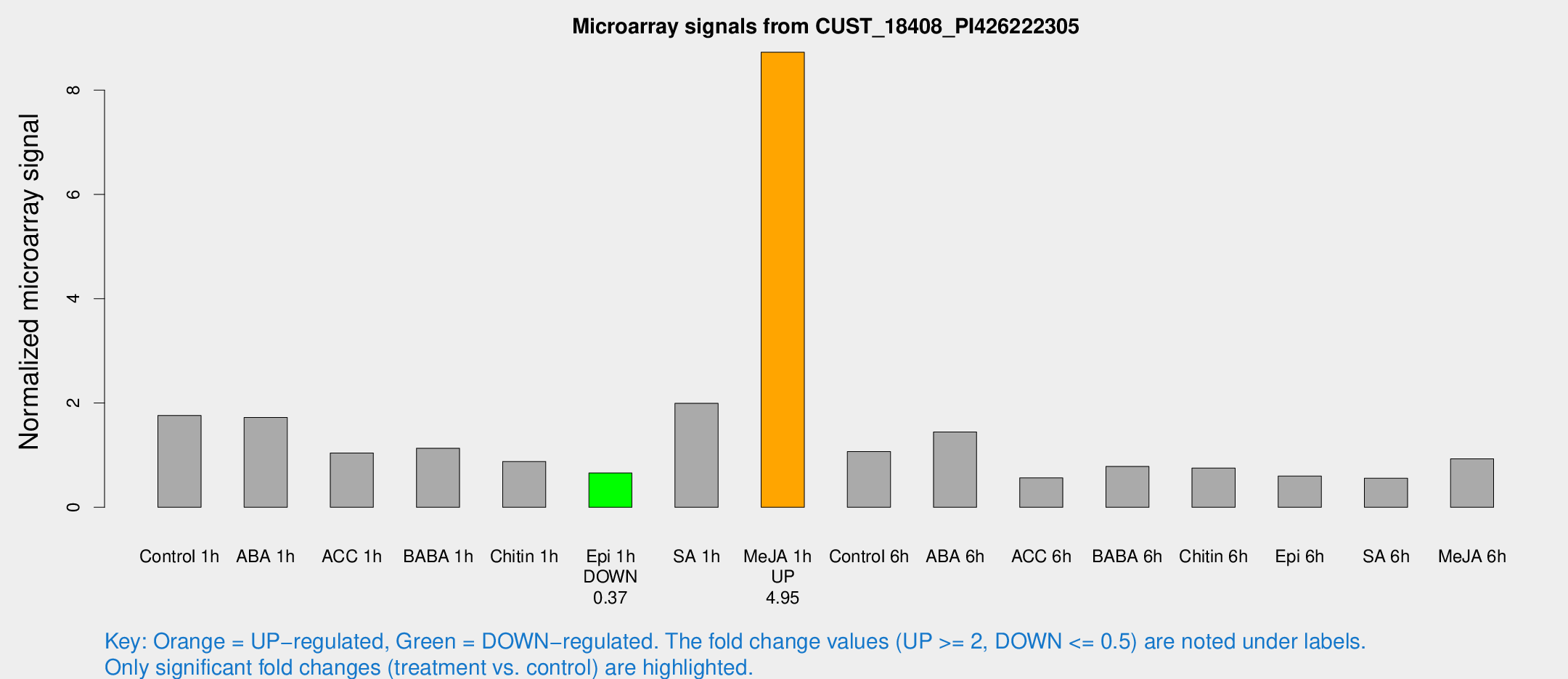
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 19.9437 | 6.36229 | 1.76206 | 0.429896 |
| ABA 1h | 18.1925 | 5.68119 | 1.72413 | 0.678951 |
| ACC 1h | 11.7061 | 3.33713 | 1.04131 | 0.342141 |
| BABA 1h | 12.6368 | 3.92948 | 1.13323 | 0.389654 |
| Chitin 1h | 8.52395 | 3.08007 | 0.878288 | 0.353847 |
| Epi 1h | 5.91912 | 3.04457 | 0.657182 | 0.344627 |
| SA 1h | 24.5526 | 9.72597 | 1.99475 | 0.898758 |
| Me-JA 1h | 74.7503 | 9.44417 | 8.72766 | 0.625135 |
| Control 6h | 12.4034 | 3.76561 | 1.06744 | 0.375644 |
| ABA 6h | 16.4114 | 3.40144 | 1.44491 | 0.319343 |
| ACC 6h | 7.02545 | 3.83903 | 0.564599 | 0.309068 |
| BABA 6h | 9.2615 | 3.55631 | 0.782576 | 0.323571 |
| Chitin 6h | 8.61514 | 3.47716 | 0.752993 | 0.330077 |
| Epi 6h | 7.09913 | 3.62281 | 0.598651 | 0.311873 |
| SA 6h | 5.69084 | 3.29801 | 0.556932 | 0.322547 |
| Me-JA 6h | 10.9432 | 4.11322 | 0.930061 | 0.361942 |
Source Transcript PGSC0003DMT400042189 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G10740.1 | +3 | 0.0 | 701 | 340/457 (74%) | alpha/beta-Hydrolases superfamily protein | chr1:3568343-3570466 REVERSE LENGTH=473 |
