Probe CUST_18019_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18019_PI426222305 | JHI_St_60k_v1 | DMT400071094 | TTACTCAAAAATGGCAACTGGTTTTCCGTGCCACCTGATCAGAATTCTTTTTTCATCAAT |
All Microarray Probes Designed to Gene DMG400027645
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18019_PI426222305 | JHI_St_60k_v1 | DMT400071094 | TTACTCAAAAATGGCAACTGGTTTTCCGTGCCACCTGATCAGAATTCTTTTTTCATCAAT |
| CUST_17920_PI426222305 | JHI_St_60k_v1 | DMT400071093 | ACCCTGTTTGAGAAATTTGGAGCCTCATAGTGATTTTTCGAACGAGAGACAAAATCAGAA |
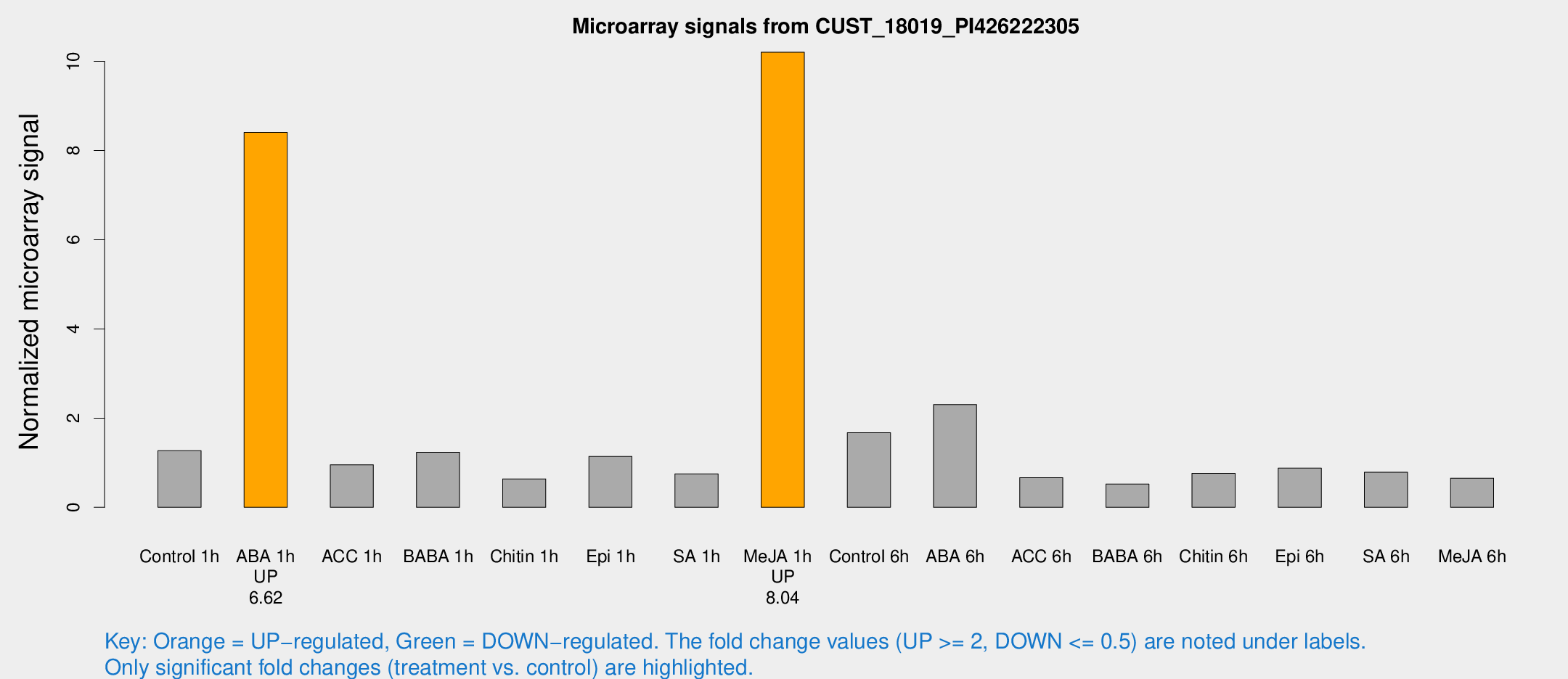
Microarray Signals from CUST_18019_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 16.9886 | 8.11583 | 1.26928 | 0.767952 |
| ABA 1h | 84.1671 | 26.9784 | 8.40695 | 2.71106 |
| ACC 1h | 10.785 | 3.32619 | 0.95367 | 0.354317 |
| BABA 1h | 12.6209 | 3.22267 | 1.23442 | 0.331586 |
| Chitin 1h | 5.82072 | 3.03057 | 0.637472 | 0.326395 |
| Epi 1h | 10.6131 | 3.02402 | 1.14206 | 0.364799 |
| SA 1h | 9.56939 | 4.34784 | 0.747879 | 0.357969 |
| Me-JA 1h | 87.894 | 12.4653 | 10.2053 | 1.02427 |
| Control 6h | 20.2541 | 7.20223 | 1.67128 | 0.594893 |
| ABA 6h | 28.8028 | 10.9828 | 2.30251 | 1.04425 |
| ACC 6h | 8.17618 | 3.81076 | 0.665724 | 0.325393 |
| BABA 6h | 6.04628 | 3.51415 | 0.522291 | 0.302759 |
| Chitin 6h | 9.35966 | 3.48554 | 0.763197 | 0.349141 |
| Epi 6h | 11.3918 | 3.77 | 0.881377 | 0.34743 |
| SA 6h | 8.98166 | 3.32382 | 0.784248 | 0.367902 |
| Me-JA 6h | 6.7731 | 3.15807 | 0.653904 | 0.312193 |
Source Transcript PGSC0003DMT400071094 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G30040.1 | +1 | 5e-123 | 365 | 190/367 (52%) | gibberellin 2-oxidase | chr1:10537769-10539570 FORWARD LENGTH=341 |
