Probe CUST_17920_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_17920_PI426222305 | JHI_St_60k_v1 | DMT400071093 | ACCCTGTTTGAGAAATTTGGAGCCTCATAGTGATTTTTCGAACGAGAGACAAAATCAGAA |
All Microarray Probes Designed to Gene DMG400027645
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_18019_PI426222305 | JHI_St_60k_v1 | DMT400071094 | TTACTCAAAAATGGCAACTGGTTTTCCGTGCCACCTGATCAGAATTCTTTTTTCATCAAT |
| CUST_17920_PI426222305 | JHI_St_60k_v1 | DMT400071093 | ACCCTGTTTGAGAAATTTGGAGCCTCATAGTGATTTTTCGAACGAGAGACAAAATCAGAA |
Microarray Signals from CUST_17920_PI426222305
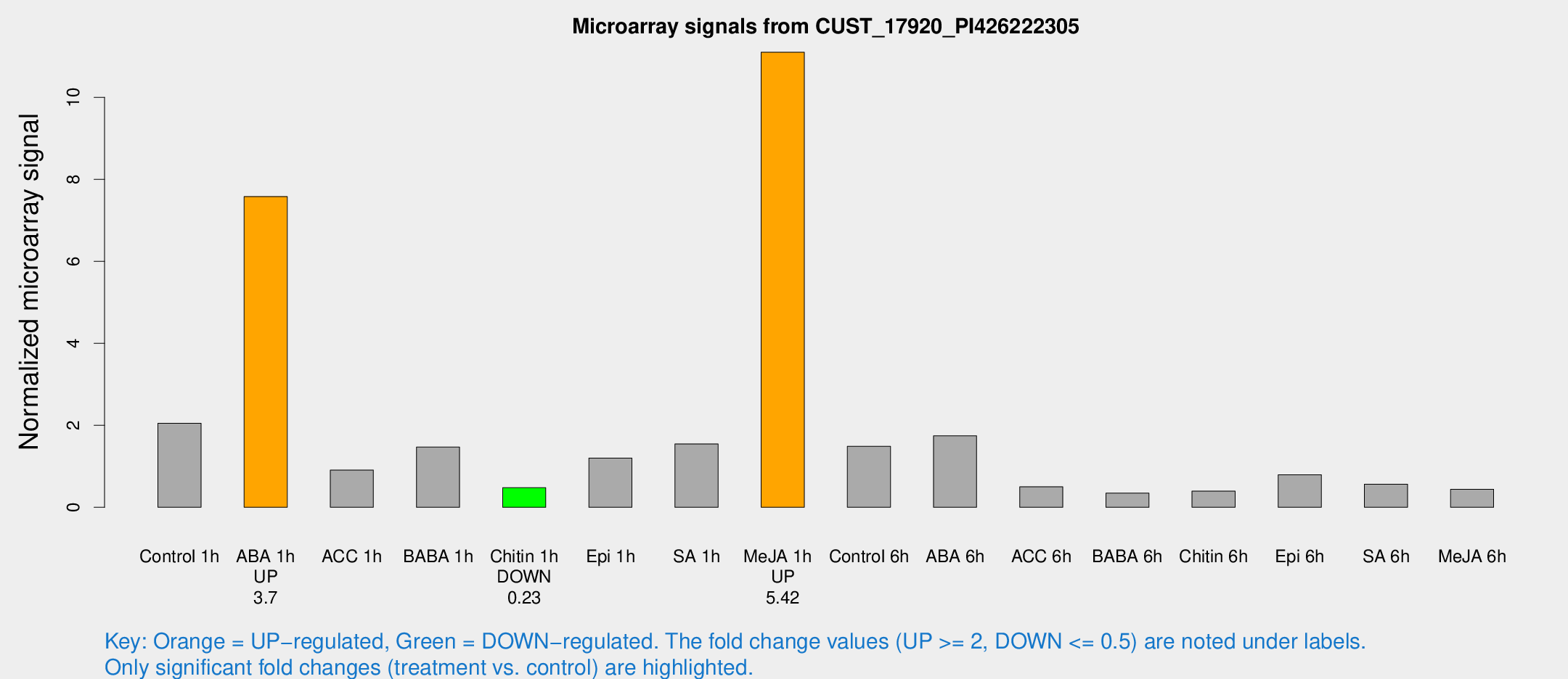
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 69.4572 | 18.4289 | 2.04863 | 0.390461 |
| ABA 1h | 219.562 | 35.3242 | 7.57977 | 1.02309 |
| ACC 1h | 35.9067 | 16.384 | 0.908055 | 0.418609 |
| BABA 1h | 45.3813 | 4.2355 | 1.46855 | 0.136037 |
| Chitin 1h | 14.7765 | 4.08049 | 0.477783 | 0.114981 |
| Epi 1h | 34.0207 | 5.3962 | 1.20208 | 0.185678 |
| SA 1h | 54.4055 | 15.1562 | 1.54652 | 0.485143 |
| Me-JA 1h | 299.495 | 54.3427 | 11.0999 | 1.45913 |
| Control 6h | 50.2937 | 10.9018 | 1.48835 | 0.241935 |
| ABA 6h | 68.8547 | 27.4332 | 1.74401 | 0.855167 |
| ACC 6h | 18.3849 | 3.91604 | 0.499284 | 0.10364 |
| BABA 6h | 14.065 | 4.42991 | 0.344833 | 0.138435 |
| Chitin 6h | 13.869 | 3.51325 | 0.395077 | 0.108995 |
| Epi 6h | 31.0502 | 8.09573 | 0.795247 | 0.207716 |
| SA 6h | 22.0022 | 8.20242 | 0.562432 | 0.41733 |
| Me-JA 6h | 15.6998 | 5.22605 | 0.440552 | 0.203455 |
Source Transcript PGSC0003DMT400071093 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G30040.1 | +2 | 4e-32 | 118 | 56/81 (69%) | gibberellin 2-oxidase | chr1:10537769-10539570 FORWARD LENGTH=341 |
