Probe CUST_16802_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_16802_PI426222305 | JHI_St_60k_v1 | DMT400069305 | TCTCATCATGACCAAAATATTCATCTCAAGTATAGTACTTGCCCAACATCACTACACTAG |
All Microarray Probes Designed to Gene DMG400026959
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_16496_PI426222305 | JHI_St_60k_v1 | DMT400069306 | GTGCATGCTACTATGATGAACAACCTAATAATGTGAAAGTGAAATCTCTTGTGGTATGAA |
| CUST_16802_PI426222305 | JHI_St_60k_v1 | DMT400069305 | TCTCATCATGACCAAAATATTCATCTCAAGTATAGTACTTGCCCAACATCACTACACTAG |
Microarray Signals from CUST_16802_PI426222305
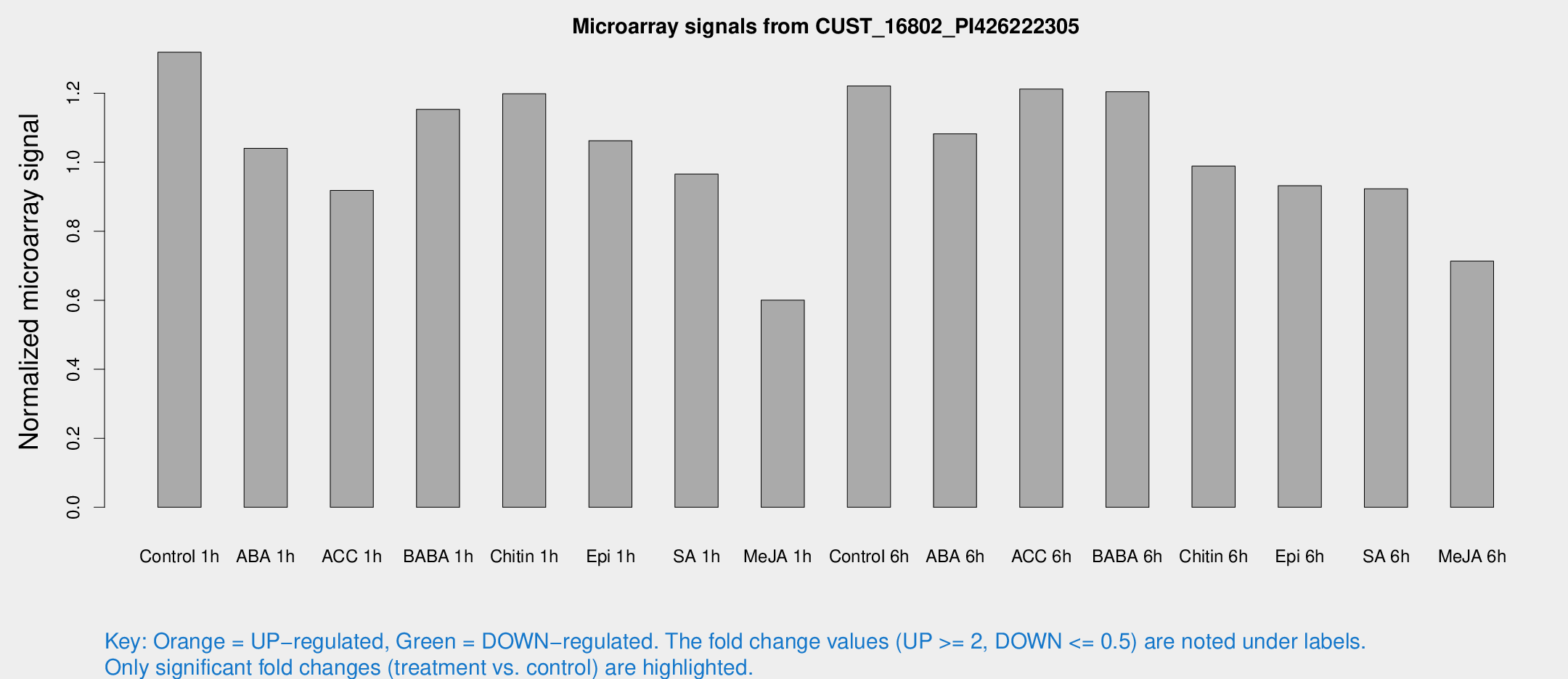
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 164.775 | 15.3316 | 1.31866 | 0.0808799 |
| ABA 1h | 114.252 | 7.34257 | 1.04016 | 0.0767801 |
| ACC 1h | 122.949 | 26.189 | 0.918181 | 0.153109 |
| BABA 1h | 138.598 | 11.176 | 1.15309 | 0.08919 |
| Chitin 1h | 138.884 | 28.6316 | 1.19808 | 0.177284 |
| Epi 1h | 119.446 | 24.9013 | 1.06217 | 0.21872 |
| SA 1h | 126.019 | 19.1922 | 0.965426 | 0.15372 |
| Me-JA 1h | 60.8092 | 5.48769 | 0.600129 | 0.064855 |
| Control 6h | 186.708 | 68.5729 | 1.22081 | 0.577795 |
| ABA 6h | 146.151 | 23.4452 | 1.08239 | 0.217235 |
| ACC 6h | 179.101 | 32.642 | 1.21183 | 0.220972 |
| BABA 6h | 168.228 | 13.8411 | 1.20406 | 0.122359 |
| Chitin 6h | 130.586 | 8.46359 | 0.9887 | 0.0641015 |
| Epi 6h | 132.455 | 17.0053 | 0.931875 | 0.0910373 |
| SA 6h | 113.131 | 7.45817 | 0.92282 | 0.113118 |
| Me-JA 6h | 90.7348 | 16.3619 | 0.71334 | 0.14429 |
Source Transcript PGSC0003DMT400069305 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G01940.3 | +1 | 9e-94 | 290 | 168/330 (51%) | C2H2-like zinc finger protein | chr2:432652-434917 FORWARD LENGTH=446 |
