Probe CUST_16725_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_16725_PI426222305 | JHI_St_60k_v1 | DMT400069497 | CGACTTCCTTAGATTCCTCGATACATAGGGAGACATTTTATAACACCATTACATGTATAT |
All Microarray Probes Designed to Gene DMG400027017
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_16725_PI426222305 | JHI_St_60k_v1 | DMT400069497 | CGACTTCCTTAGATTCCTCGATACATAGGGAGACATTTTATAACACCATTACATGTATAT |
| CUST_16700_PI426222305 | JHI_St_60k_v1 | DMT400069498 | TTTCCAGATTATGAGTCTAGTGACAAGAAGGATCTACTTGAGCTTGGTGAATTTGAGTGG |
Microarray Signals from CUST_16725_PI426222305
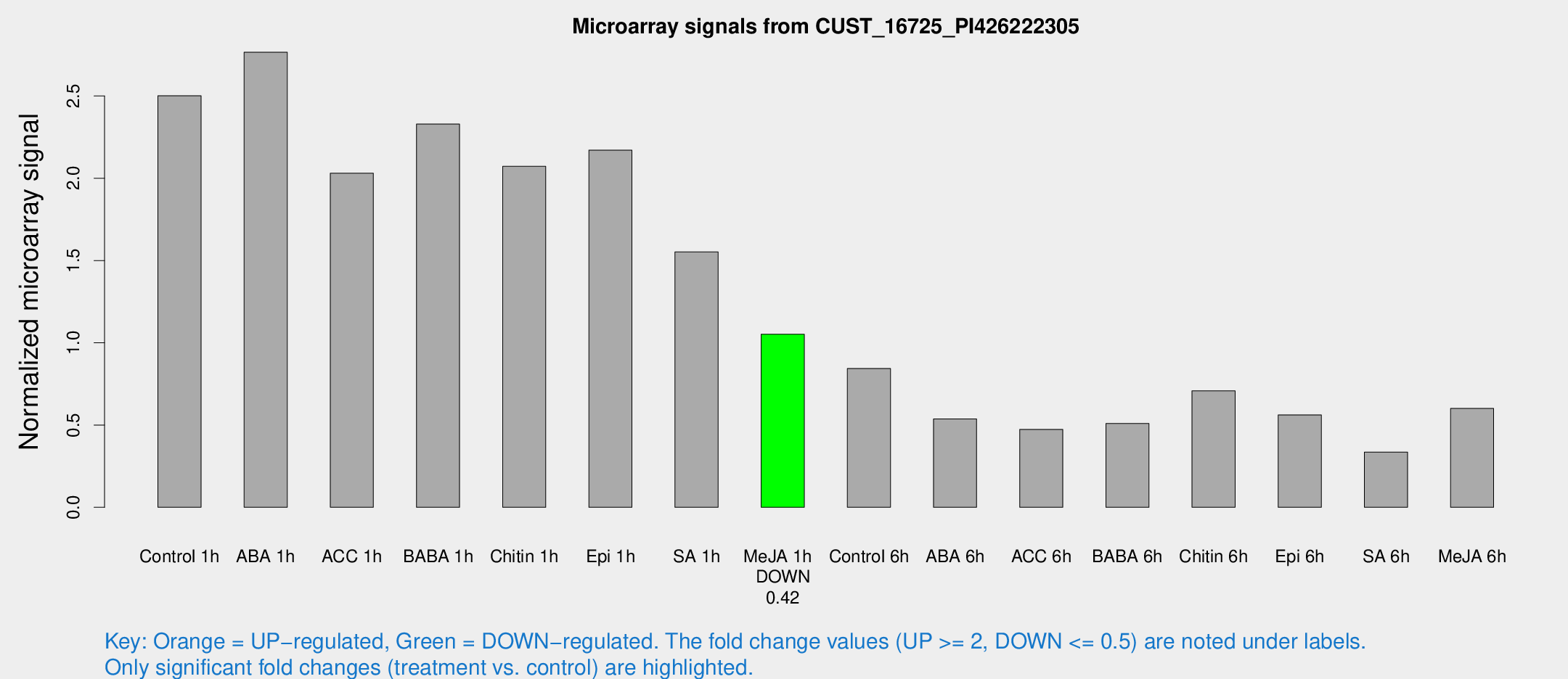
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 871.653 | 121.392 | 2.50135 | 0.19841 |
| ABA 1h | 846.801 | 86.2352 | 2.76608 | 0.479248 |
| ACC 1h | 726.527 | 94.1338 | 2.0304 | 0.120655 |
| BABA 1h | 803.389 | 156.065 | 2.32952 | 0.26939 |
| Chitin 1h | 638.974 | 37.0933 | 2.07222 | 0.144857 |
| Epi 1h | 658.213 | 102.372 | 2.17079 | 0.339614 |
| SA 1h | 548.924 | 41.8845 | 1.55198 | 0.0900593 |
| Me-JA 1h | 299.017 | 40.5943 | 1.05155 | 0.0617843 |
| Control 6h | 313.314 | 86.1793 | 0.843909 | 0.189413 |
| ABA 6h | 198.365 | 23.3172 | 0.537142 | 0.0514539 |
| ACC 6h | 189.884 | 28.7795 | 0.473421 | 0.028867 |
| BABA 6h | 203.001 | 40.4607 | 0.509274 | 0.0920903 |
| Chitin 6h | 261.894 | 32.7872 | 0.707905 | 0.0759697 |
| Epi 6h | 225.927 | 47.6296 | 0.561792 | 0.128067 |
| SA 6h | 139.368 | 65.7585 | 0.335708 | 0.145405 |
| Me-JA 6h | 215.124 | 44.1137 | 0.600778 | 0.0904553 |
Source Transcript PGSC0003DMT400069497 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G06040.1 | +2 | 1e-29 | 119 | 63/108 (58%) | B-box zinc finger family protein | chr1:1828662-1829659 REVERSE LENGTH=248 |
