Probe CUST_16700_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_16700_PI426222305 | JHI_St_60k_v1 | DMT400069498 | TTTCCAGATTATGAGTCTAGTGACAAGAAGGATCTACTTGAGCTTGGTGAATTTGAGTGG |
All Microarray Probes Designed to Gene DMG400027017
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_16725_PI426222305 | JHI_St_60k_v1 | DMT400069497 | CGACTTCCTTAGATTCCTCGATACATAGGGAGACATTTTATAACACCATTACATGTATAT |
| CUST_16700_PI426222305 | JHI_St_60k_v1 | DMT400069498 | TTTCCAGATTATGAGTCTAGTGACAAGAAGGATCTACTTGAGCTTGGTGAATTTGAGTGG |
Microarray Signals from CUST_16700_PI426222305
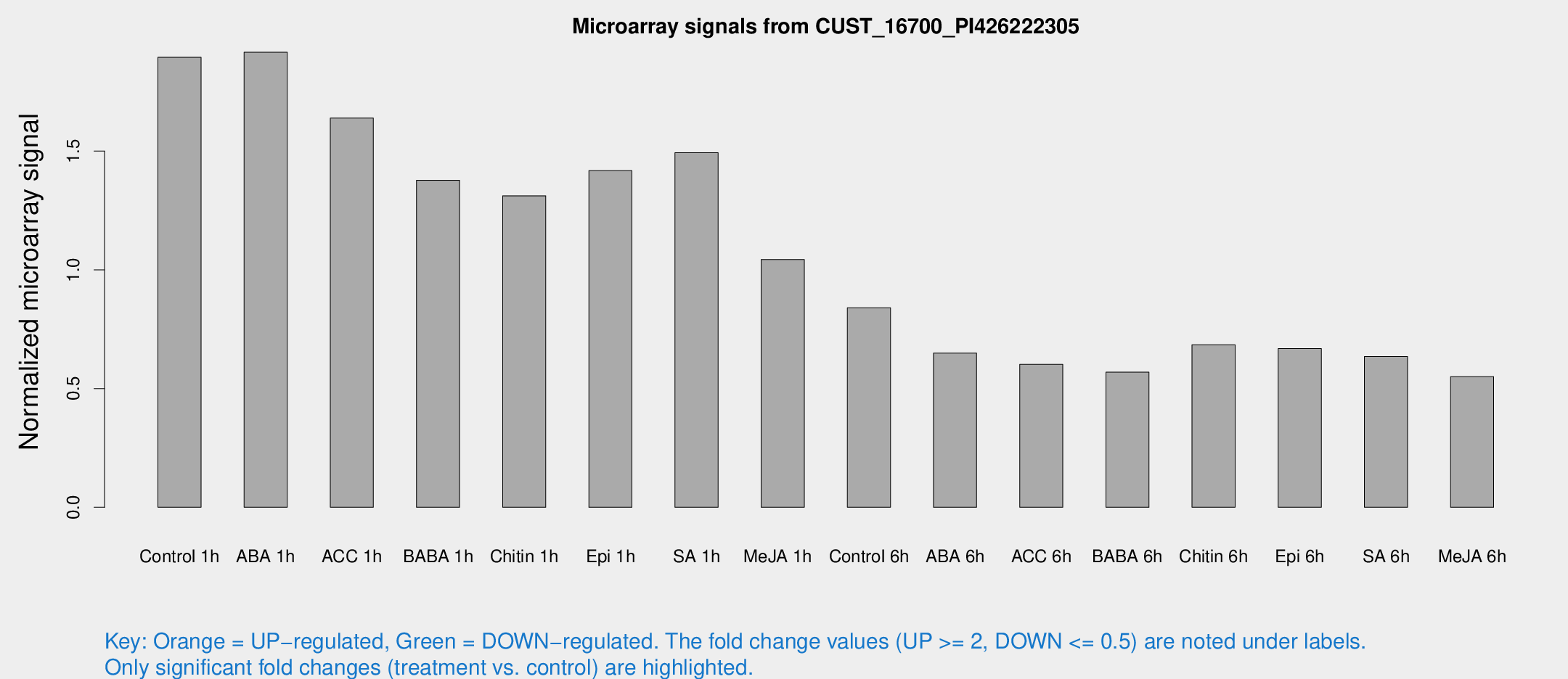
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 21641.4 | 4096.15 | 1.89528 | 0.223933 |
| ABA 1h | 18918.2 | 1869.14 | 1.91662 | 0.121299 |
| ACC 1h | 18795.7 | 1889.45 | 1.63962 | 0.0946641 |
| BABA 1h | 15717.9 | 3713.15 | 1.37725 | 0.24366 |
| Chitin 1h | 13318.9 | 1899.05 | 1.31189 | 0.105534 |
| Epi 1h | 13601.5 | 820.467 | 1.41796 | 0.0818664 |
| SA 1h | 17198 | 2187.12 | 1.49346 | 0.107703 |
| Me-JA 1h | 9511.04 | 992.136 | 1.04376 | 0.0602624 |
| Control 6h | 9728.17 | 2063.02 | 0.84014 | 0.131263 |
| ABA 6h | 7700.69 | 756.044 | 0.649709 | 0.0582476 |
| ACC 6h | 7835.78 | 1296.19 | 0.601951 | 0.0347547 |
| BABA 6h | 7130.96 | 870.029 | 0.568949 | 0.0646085 |
| Chitin 6h | 8151.22 | 926.58 | 0.684846 | 0.0645065 |
| Epi 6h | 8363.56 | 589.319 | 0.66841 | 0.0873204 |
| SA 6h | 7517.61 | 1953.68 | 0.634835 | 0.119103 |
| Me-JA 6h | 6203.54 | 968.393 | 0.550348 | 0.0406858 |
Source Transcript PGSC0003DMT400069498 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G06040.1 | +3 | 6e-86 | 267 | 148/248 (60%) | B-box zinc finger family protein | chr1:1828662-1829659 REVERSE LENGTH=248 |
