Probe CUST_15624_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_15624_PI426222305 | JHI_St_60k_v1 | DMT400004788 | TGCTTTGGTTACTTACAGCCACGGCCCATGTCATCAATAAAATTGGATGCCCCAATCTGA |
All Microarray Probes Designed to Gene DMG401001903
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_15605_PI426222305 | JHI_St_60k_v1 | DMT400004790 | GTCAGTGTGTGTACTATTTTGTCAGACTAATAGGATAACAAAGTTTGAAGCTAAGTGTCA |
| CUST_15624_PI426222305 | JHI_St_60k_v1 | DMT400004788 | TGCTTTGGTTACTTACAGCCACGGCCCATGTCATCAATAAAATTGGATGCCCCAATCTGA |
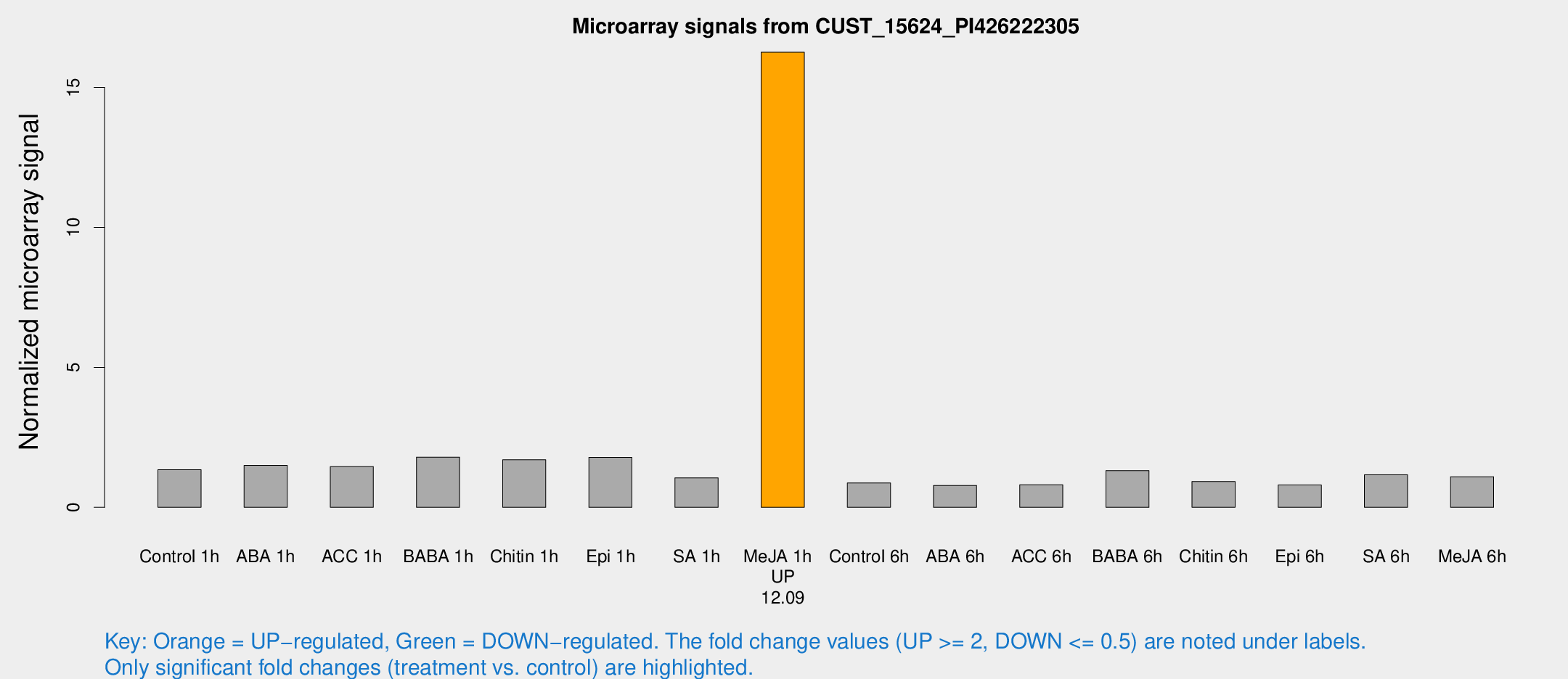
Microarray Signals from CUST_15624_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 10.4769 | 3.23267 | 1.34514 | 0.473296 |
| ABA 1h | 10.4389 | 3.19327 | 1.5001 | 0.542352 |
| ACC 1h | 13.5905 | 6.55624 | 1.45405 | 0.688613 |
| BABA 1h | 13.4493 | 3.4971 | 1.79236 | 0.534252 |
| Chitin 1h | 12.2948 | 3.6549 | 1.69655 | 0.754261 |
| Epi 1h | 12.25 | 3.271 | 1.78338 | 0.613935 |
| SA 1h | 8.08733 | 3.24672 | 1.04982 | 0.449187 |
| Me-JA 1h | 105.739 | 28.222 | 16.2587 | 3.64042 |
| Control 6h | 6.40656 | 3.4708 | 0.86723 | 0.473817 |
| ABA 6h | 6.08898 | 3.5292 | 0.781112 | 0.452458 |
| ACC 6h | 6.92342 | 4.10109 | 0.806035 | 0.466778 |
| BABA 6h | 13.5264 | 6.70974 | 1.31135 | 0.693512 |
| Chitin 6h | 7.28603 | 3.76686 | 0.924446 | 0.4883 |
| Epi 6h | 6.66165 | 3.90667 | 0.79795 | 0.462726 |
| SA 6h | 8.74158 | 3.63275 | 1.16087 | 0.521662 |
| Me-JA 6h | 8.68869 | 3.50868 | 1.09344 | 0.523286 |
Source Transcript PGSC0003DMT400004788 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G31940.1 | +1 | 2e-70 | 231 | 134/304 (44%) | cytochrome P450, family 82, subfamily C, polypeptide 4 | chr4:15452040-15453966 FORWARD LENGTH=524 |
