Probe CUST_15605_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_15605_PI426222305 | JHI_St_60k_v1 | DMT400004790 | GTCAGTGTGTGTACTATTTTGTCAGACTAATAGGATAACAAAGTTTGAAGCTAAGTGTCA |
All Microarray Probes Designed to Gene DMG401001903
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_15605_PI426222305 | JHI_St_60k_v1 | DMT400004790 | GTCAGTGTGTGTACTATTTTGTCAGACTAATAGGATAACAAAGTTTGAAGCTAAGTGTCA |
| CUST_15624_PI426222305 | JHI_St_60k_v1 | DMT400004788 | TGCTTTGGTTACTTACAGCCACGGCCCATGTCATCAATAAAATTGGATGCCCCAATCTGA |
Microarray Signals from CUST_15605_PI426222305
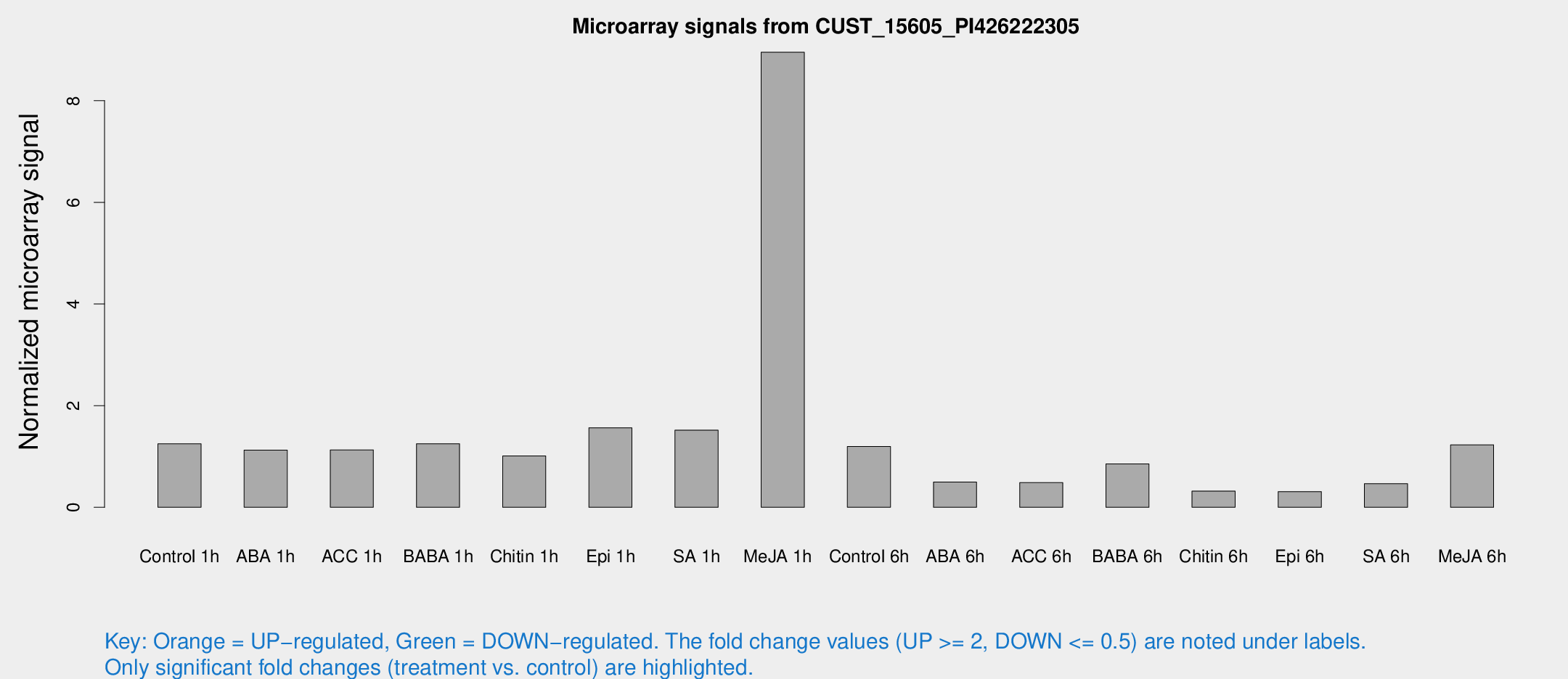
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 55.3944 | 15.6762 | 1.24882 | 0.26108 |
| ABA 1h | 42.3006 | 7.39687 | 1.12615 | 0.229246 |
| ACC 1h | 57.8745 | 26.0285 | 1.12845 | 0.48551 |
| BABA 1h | 53.5264 | 14.5809 | 1.25156 | 0.251354 |
| Chitin 1h | 45.1243 | 16.7594 | 1.00977 | 0.530088 |
| Epi 1h | 56.9973 | 7.84444 | 1.56507 | 0.217272 |
| SA 1h | 66.8199 | 12.8891 | 1.51861 | 0.360164 |
| Me-JA 1h | 334.3 | 103.146 | 8.95434 | 2.24582 |
| Control 6h | 58.6754 | 22.8168 | 1.19622 | 0.463848 |
| ABA 6h | 23.686 | 6.02751 | 0.495647 | 0.140671 |
| ACC 6h | 31.0648 | 16.4186 | 0.485127 | 0.396437 |
| BABA 6h | 55.1848 | 31.1525 | 0.854427 | 0.620174 |
| Chitin 6h | 15.4561 | 5.01571 | 0.319731 | 0.114555 |
| Epi 6h | 15.7028 | 4.18354 | 0.308649 | 0.127501 |
| SA 6h | 19.6476 | 3.91822 | 0.463806 | 0.0938394 |
| Me-JA 6h | 68.8982 | 28.3976 | 1.22973 | 1.35626 |
Source Transcript PGSC0003DMT400004790 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G31940.1 | +1 | 5e-163 | 479 | 249/511 (49%) | cytochrome P450, family 82, subfamily C, polypeptide 4 | chr4:15452040-15453966 FORWARD LENGTH=524 |
