Probe CUST_15149_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_15149_PI426222305 | JHI_St_60k_v1 | DMT400057198 | TTTGCTTAGCCAGTACGGTGACATCTCATTCCTTTGTTTTATCATTTATTACCTGACGAC |
All Microarray Probes Designed to Gene DMG400022217
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14956_PI426222305 | JHI_St_60k_v1 | DMT400057197 | GCATCATAATGAAGTCCATCATAAATCAGCATTACTCTTTCCTGATAGTTCTTTCCCTGT |
| CUST_15149_PI426222305 | JHI_St_60k_v1 | DMT400057198 | TTTGCTTAGCCAGTACGGTGACATCTCATTCCTTTGTTTTATCATTTATTACCTGACGAC |
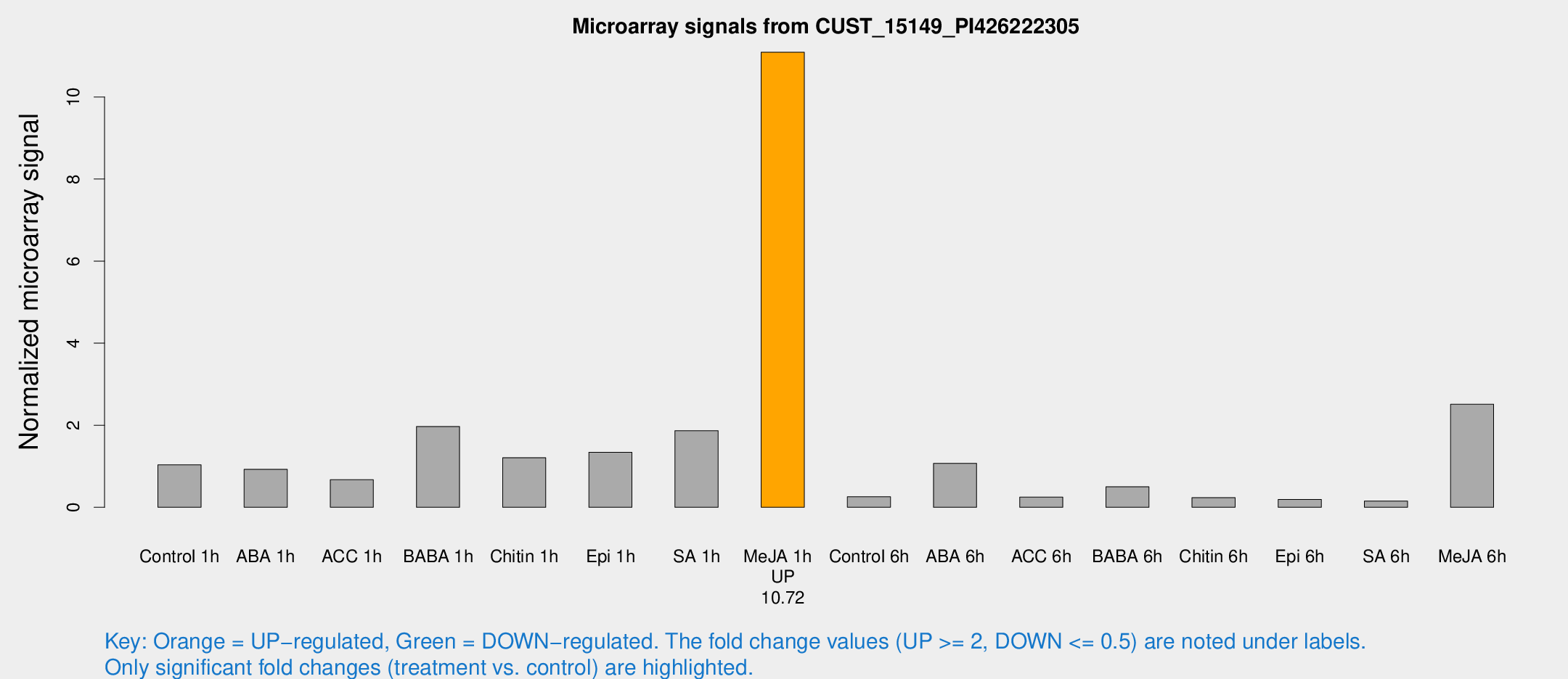
Microarray Signals from CUST_15149_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 62.3674 | 5.16706 | 1.03475 | 0.0968516 |
| ABA 1h | 60.5966 | 23.0547 | 0.926094 | 0.472366 |
| ACC 1h | 51.2614 | 24.4117 | 0.671152 | 0.31496 |
| BABA 1h | 114.662 | 10.7301 | 1.96982 | 0.133466 |
| Chitin 1h | 66.8484 | 11.4556 | 1.20842 | 0.224606 |
| Epi 1h | 80.0834 | 27.2957 | 1.34047 | 0.585966 |
| SA 1h | 125.561 | 35.8571 | 1.86625 | 0.705809 |
| Me-JA 1h | 547.012 | 52.6673 | 11.0909 | 0.644752 |
| Control 6h | 33.2463 | 26.7335 | 0.256953 | 0.368655 |
| ABA 6h | 69.4559 | 9.36422 | 1.07188 | 0.0882712 |
| ACC 6h | 19.0546 | 6.78421 | 0.247213 | 0.118101 |
| BABA 6h | 65.9337 | 50.9139 | 0.499562 | 0.691003 |
| Chitin 6h | 15.5407 | 4.33972 | 0.235193 | 0.0724213 |
| Epi 6h | 15.1921 | 6.5189 | 0.190285 | 0.104713 |
| SA 6h | 9.44882 | 4.06145 | 0.153091 | 0.0718462 |
| Me-JA 6h | 150.109 | 9.51486 | 2.51199 | 0.1589 |
Source Transcript PGSC0003DMT400057198 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | None | - | - | - | - | - |
