Probe CUST_14956_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14956_PI426222305 | JHI_St_60k_v1 | DMT400057197 | GCATCATAATGAAGTCCATCATAAATCAGCATTACTCTTTCCTGATAGTTCTTTCCCTGT |
All Microarray Probes Designed to Gene DMG400022217
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14956_PI426222305 | JHI_St_60k_v1 | DMT400057197 | GCATCATAATGAAGTCCATCATAAATCAGCATTACTCTTTCCTGATAGTTCTTTCCCTGT |
| CUST_15149_PI426222305 | JHI_St_60k_v1 | DMT400057198 | TTTGCTTAGCCAGTACGGTGACATCTCATTCCTTTGTTTTATCATTTATTACCTGACGAC |
Microarray Signals from CUST_14956_PI426222305
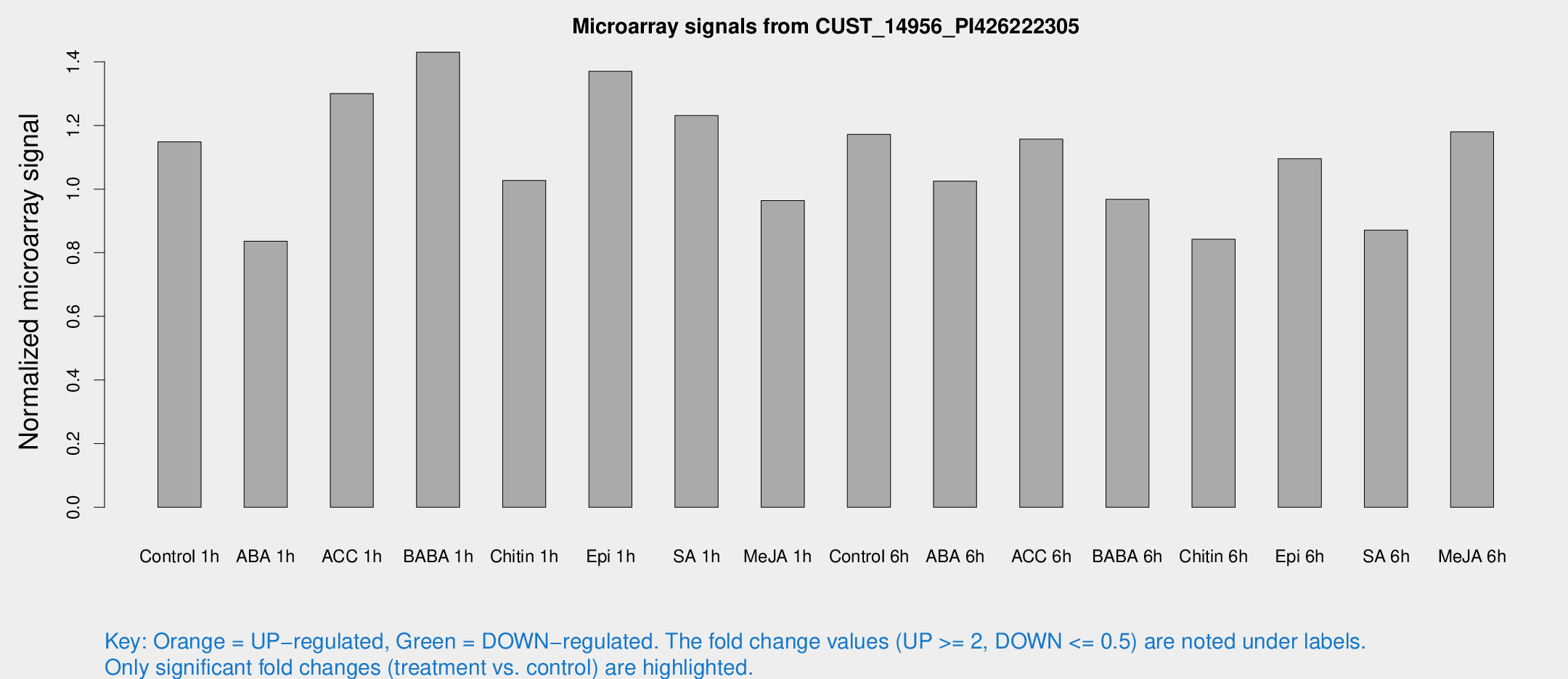
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 9.81277 | 3.73655 | 1.14834 | 0.481767 |
| ABA 1h | 6.09249 | 3.53481 | 0.836098 | 0.484896 |
| ACC 1h | 11.7784 | 4.81908 | 1.30035 | 0.605406 |
| BABA 1h | 12.4808 | 4.00203 | 1.43 | 0.550135 |
| Chitin 1h | 7.97664 | 3.68375 | 1.02705 | 0.510741 |
| Epi 1h | 10.2057 | 3.60108 | 1.36999 | 0.546023 |
| SA 1h | 10.5569 | 3.73766 | 1.23101 | 0.443352 |
| Me-JA 1h | 6.53153 | 3.8186 | 0.96383 | 0.558858 |
| Control 6h | 10.8154 | 3.93398 | 1.17204 | 0.546284 |
| ABA 6h | 9.19777 | 4.02666 | 1.02481 | 0.469391 |
| ACC 6h | 10.9829 | 4.49216 | 1.15693 | 0.468741 |
| BABA 6h | 9.16745 | 4.32433 | 0.967616 | 0.484239 |
| Chitin 6h | 7.3988 | 4.29795 | 0.842607 | 0.488552 |
| Epi 6h | 10.9165 | 4.62505 | 1.09531 | 0.522839 |
| SA 6h | 7.09759 | 4.11131 | 0.871238 | 0.504517 |
| Me-JA 6h | 10.148 | 3.90142 | 1.17974 | 0.501562 |
Source Transcript PGSC0003DMT400057197 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G20390.1 | +3 | 5e-55 | 189 | 101/172 (59%) | endoribonuclease L-PSP family protein | chr3:7110227-7111695 REVERSE LENGTH=187 |
