Probe CUST_14242_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14242_PI426222305 | JHI_St_60k_v1 | DMT400060396 | GGCGGGTTGTGTTACATTTGTATGGATTAAAATCGGCGTTCAACTTAAAATCTCTTCTTT |
All Microarray Probes Designed to Gene DMG400023490
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14150_PI426222305 | JHI_St_60k_v1 | DMT400060395 | GGCGGGTTGTGTTACATTTGTATGGATTAAAATCGGCGTTCAACTTAAAATCTCTTCTTT |
| CUST_14242_PI426222305 | JHI_St_60k_v1 | DMT400060396 | GGCGGGTTGTGTTACATTTGTATGGATTAAAATCGGCGTTCAACTTAAAATCTCTTCTTT |
Microarray Signals from CUST_14242_PI426222305
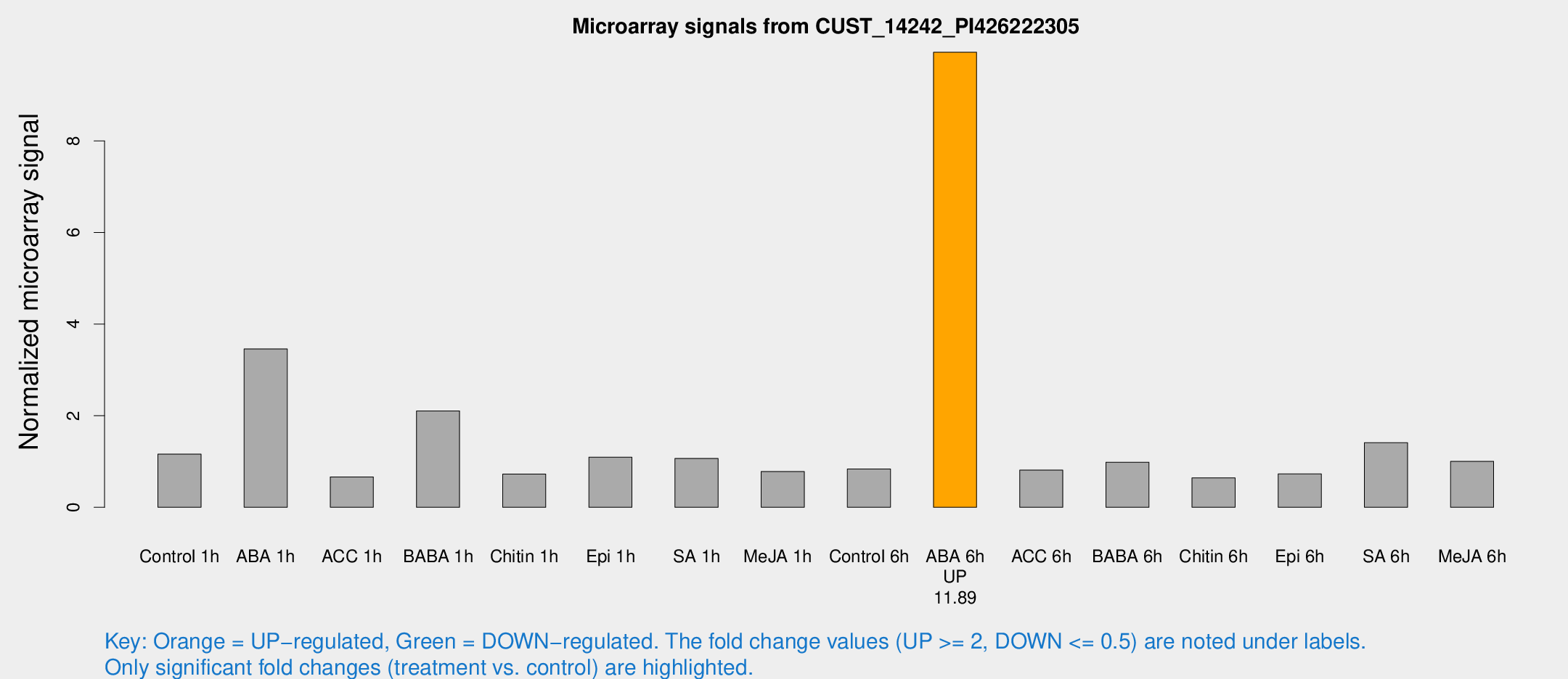
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 36.8888 | 6.70343 | 1.16077 | 0.2505 |
| ABA 1h | 95.2278 | 13.431 | 3.45731 | 0.528081 |
| ACC 1h | 22.5981 | 6.7101 | 0.664454 | 0.169593 |
| BABA 1h | 64.4754 | 12.5472 | 2.10506 | 0.662665 |
| Chitin 1h | 26.1495 | 10.3162 | 0.724597 | 0.566263 |
| Epi 1h | 29.398 | 3.94237 | 1.09447 | 0.153614 |
| SA 1h | 33.6814 | 4.20724 | 1.06681 | 0.144851 |
| Me-JA 1h | 21.4865 | 6.73549 | 0.779746 | 0.321893 |
| Control 6h | 26.18 | 4.22653 | 0.835545 | 0.144699 |
| ABA 6h | 326.562 | 33.7768 | 9.93756 | 0.587647 |
| ACC 6h | 28.6158 | 4.84501 | 0.811379 | 0.135971 |
| BABA 6h | 33.731 | 4.75982 | 0.984569 | 0.138237 |
| Chitin 6h | 21.4649 | 4.54923 | 0.643863 | 0.142059 |
| Epi 6h | 25.6252 | 4.40652 | 0.72877 | 0.14967 |
| SA 6h | 42.7252 | 4.62866 | 1.41008 | 0.153506 |
| Me-JA 6h | 31.0915 | 4.48002 | 1.00308 | 0.147668 |
Source Transcript PGSC0003DMT400060396 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G36380.1 | +3 | 0.0 | 1969 | 1017/1454 (70%) | pleiotropic drug resistance 6 | chr2:15257583-15263627 FORWARD LENGTH=1453 |
