Probe CUST_14150_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14150_PI426222305 | JHI_St_60k_v1 | DMT400060395 | GGCGGGTTGTGTTACATTTGTATGGATTAAAATCGGCGTTCAACTTAAAATCTCTTCTTT |
All Microarray Probes Designed to Gene DMG400023490
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_14150_PI426222305 | JHI_St_60k_v1 | DMT400060395 | GGCGGGTTGTGTTACATTTGTATGGATTAAAATCGGCGTTCAACTTAAAATCTCTTCTTT |
| CUST_14242_PI426222305 | JHI_St_60k_v1 | DMT400060396 | GGCGGGTTGTGTTACATTTGTATGGATTAAAATCGGCGTTCAACTTAAAATCTCTTCTTT |
Microarray Signals from CUST_14150_PI426222305
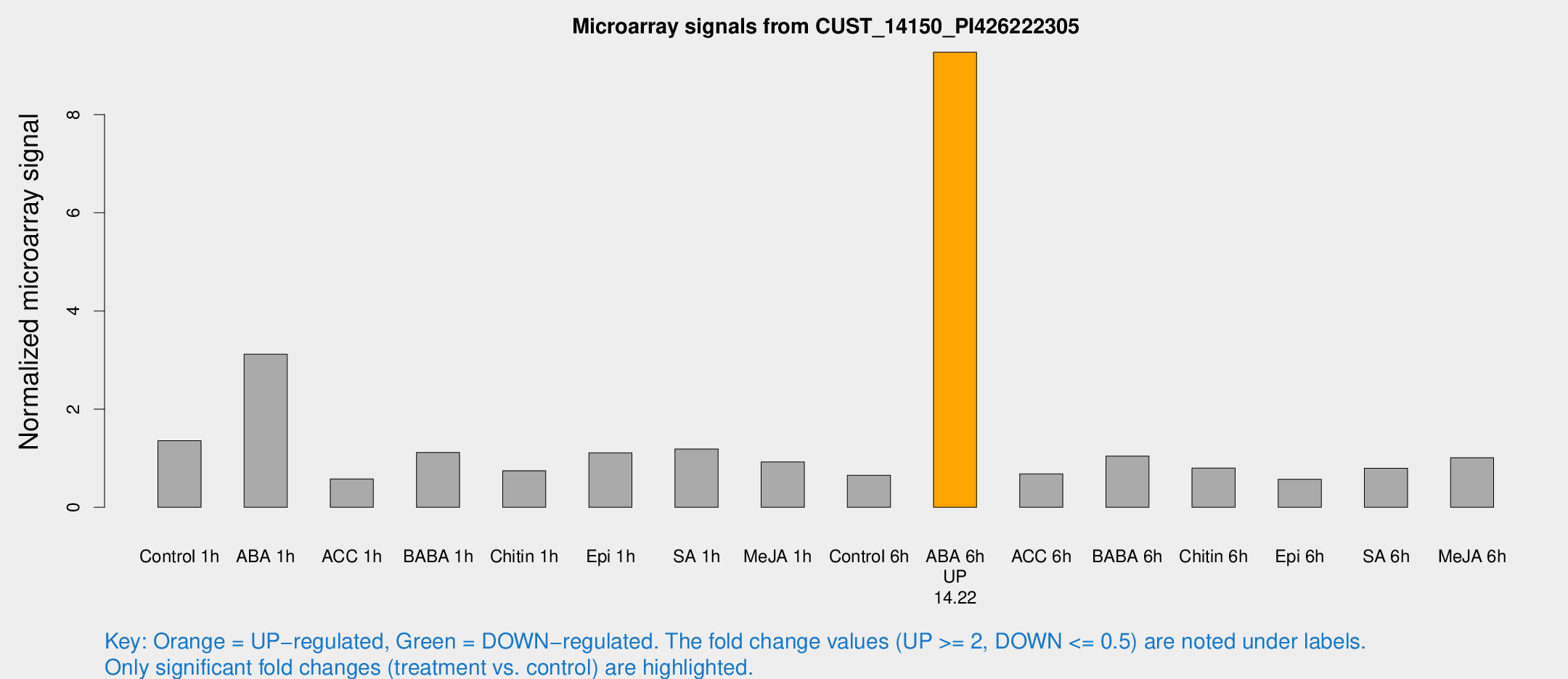
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 41.1731 | 4.28535 | 1.3574 | 0.140642 |
| ABA 1h | 87.9866 | 21.7227 | 3.11749 | 0.742936 |
| ACC 1h | 19.4226 | 5.2973 | 0.578298 | 0.190828 |
| BABA 1h | 32.4101 | 4.11616 | 1.11751 | 0.142582 |
| Chitin 1h | 22.5993 | 7.30724 | 0.745002 | 0.216537 |
| Epi 1h | 33.7888 | 11.5307 | 1.1099 | 0.534464 |
| SA 1h | 37.3457 | 5.23455 | 1.18564 | 0.257768 |
| Me-JA 1h | 24.9252 | 7.12634 | 0.923627 | 0.433858 |
| Control 6h | 22.1724 | 6.64753 | 0.651758 | 0.199782 |
| ABA 6h | 302.732 | 48.0729 | 9.27001 | 0.920084 |
| ACC 6h | 25.5227 | 7.88778 | 0.679823 | 0.131446 |
| BABA 6h | 35.1851 | 4.45694 | 1.04374 | 0.133321 |
| Chitin 6h | 25.6771 | 4.20481 | 0.800367 | 0.132808 |
| Epi 6h | 21.6989 | 6.23838 | 0.57158 | 0.180861 |
| SA 6h | 26.5335 | 7.77036 | 0.794588 | 0.209041 |
| Me-JA 6h | 30.255 | 3.89998 | 1.00816 | 0.131471 |
Source Transcript PGSC0003DMT400060395 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G36380.1 | +3 | 0.0 | 1514 | 781/1102 (71%) | pleiotropic drug resistance 6 | chr2:15257583-15263627 FORWARD LENGTH=1453 |
