Probe CUST_13949_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_13949_PI426222305 | JHI_St_60k_v1 | DMT400061440 | CACCCCTTTTCAAGGGATGTGATATGTAAAATTTTATGCTCTACTGAAAATTTTAGCACC |
All Microarray Probes Designed to Gene DMG400023912
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_13908_PI426222305 | JHI_St_60k_v1 | DMT400061439 | CACCCCTTTTCAAGGGATGTGATATGTAAAATTTTATGCTCTACTGAAAATTTTAGCACC |
| CUST_13949_PI426222305 | JHI_St_60k_v1 | DMT400061440 | CACCCCTTTTCAAGGGATGTGATATGTAAAATTTTATGCTCTACTGAAAATTTTAGCACC |
Microarray Signals from CUST_13949_PI426222305
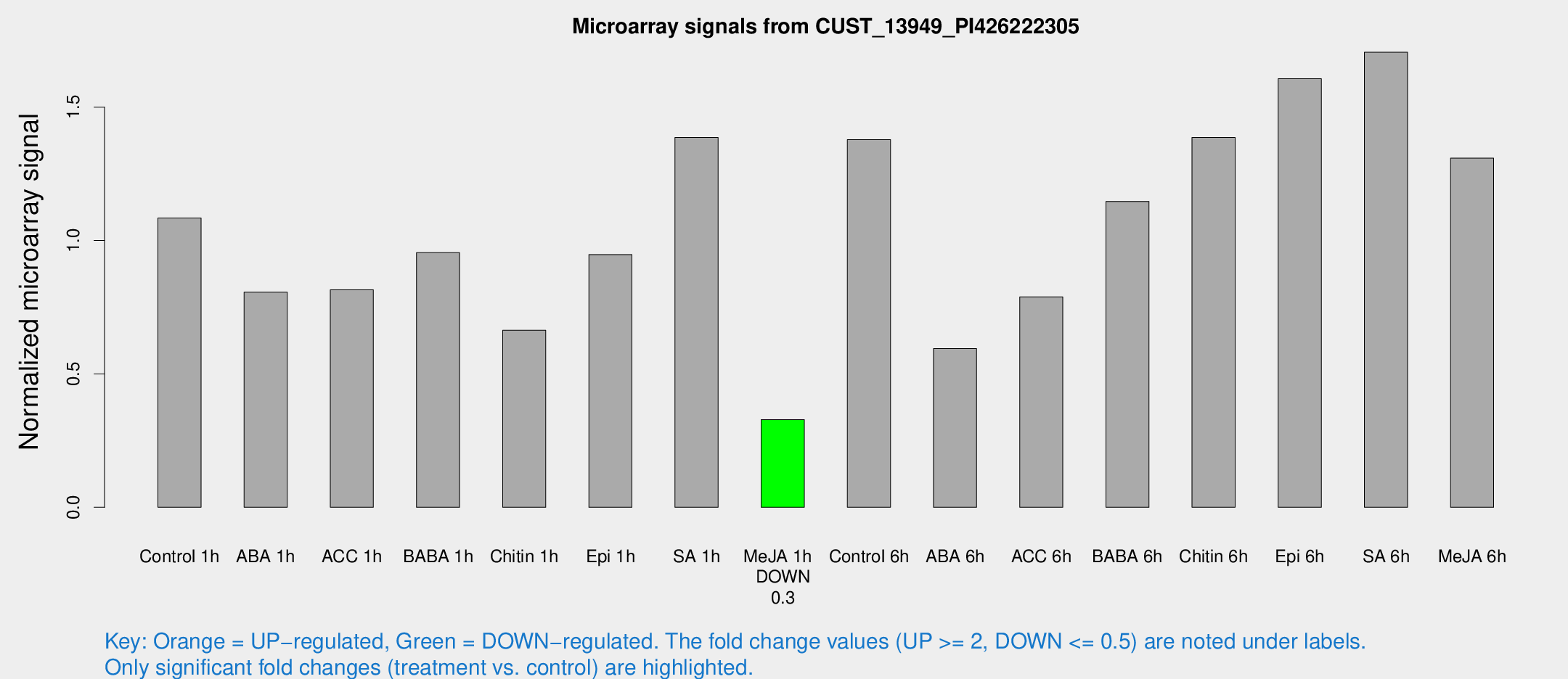
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 91.5114 | 32.8193 | 1.08431 | 0.315359 |
| ABA 1h | 54.0861 | 5.07426 | 0.806539 | 0.0739084 |
| ACC 1h | 78.337 | 29.3596 | 0.815546 | 0.421364 |
| BABA 1h | 71.9073 | 13.2564 | 0.954594 | 0.0989762 |
| Chitin 1h | 45.3856 | 4.81347 | 0.663779 | 0.0616263 |
| Epi 1h | 62.7131 | 7.83847 | 0.9476 | 0.115569 |
| SA 1h | 107.67 | 8.58984 | 1.38628 | 0.0908422 |
| Me-JA 1h | 20.6831 | 3.60223 | 0.328628 | 0.0587191 |
| Control 6h | 114.264 | 31.0074 | 1.37832 | 0.334104 |
| ABA 6h | 49.7375 | 10.0157 | 0.595022 | 0.100273 |
| ACC 6h | 69.0811 | 8.31103 | 0.788758 | 0.0760805 |
| BABA 6h | 98.4936 | 13.4125 | 1.14669 | 0.128573 |
| Chitin 6h | 112.994 | 15.0755 | 1.38661 | 0.177856 |
| Epi 6h | 136.742 | 8.86375 | 1.60685 | 0.209428 |
| SA 6h | 126.863 | 8.17192 | 1.70598 | 0.220187 |
| Me-JA 6h | 104.985 | 26.8866 | 1.30864 | 0.244686 |
Source Transcript PGSC0003DMT400061440 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G07710.1 | +3 | 1e-156 | 462 | 268/430 (62%) | Ankyrin repeat family protein | chr1:2386275-2387986 REVERSE LENGTH=543 |
