Probe CUST_13908_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_13908_PI426222305 | JHI_St_60k_v1 | DMT400061439 | CACCCCTTTTCAAGGGATGTGATATGTAAAATTTTATGCTCTACTGAAAATTTTAGCACC |
All Microarray Probes Designed to Gene DMG400023912
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_13908_PI426222305 | JHI_St_60k_v1 | DMT400061439 | CACCCCTTTTCAAGGGATGTGATATGTAAAATTTTATGCTCTACTGAAAATTTTAGCACC |
| CUST_13949_PI426222305 | JHI_St_60k_v1 | DMT400061440 | CACCCCTTTTCAAGGGATGTGATATGTAAAATTTTATGCTCTACTGAAAATTTTAGCACC |
Microarray Signals from CUST_13908_PI426222305
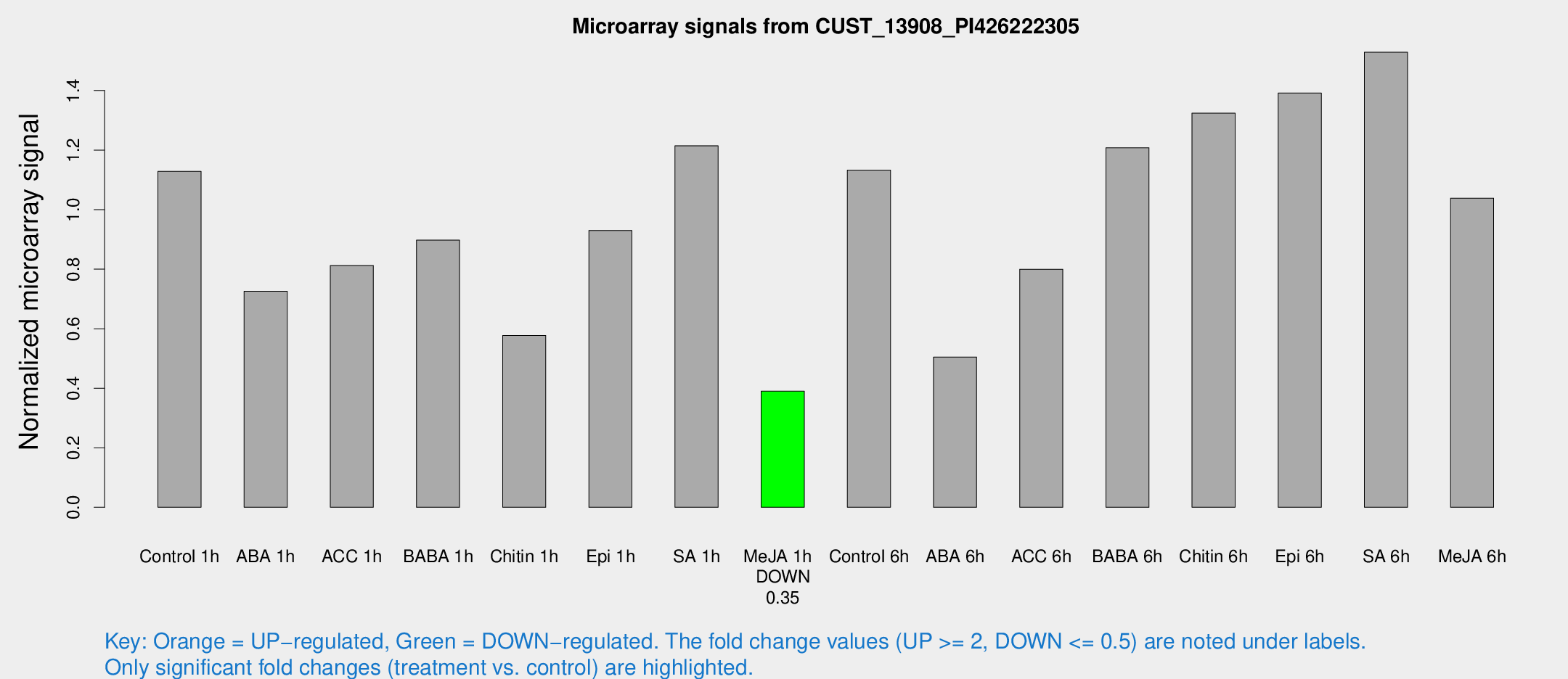
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 102.742 | 28.9087 | 1.12877 | 0.233026 |
| ABA 1h | 55.5991 | 7.94121 | 0.725582 | 0.0764966 |
| ACC 1h | 79.8148 | 25.2283 | 0.812346 | 0.258637 |
| BABA 1h | 78.1563 | 17.7244 | 0.897488 | 0.149055 |
| Chitin 1h | 44.7727 | 6.25248 | 0.577384 | 0.0578298 |
| Epi 1h | 72.2769 | 18.1086 | 0.930031 | 0.204688 |
| SA 1h | 109.121 | 20.5012 | 1.21471 | 0.164547 |
| Me-JA 1h | 27.3533 | 4.04015 | 0.390056 | 0.0578721 |
| Control 6h | 101.299 | 21.6897 | 1.13265 | 0.16353 |
| ABA 6h | 47.4341 | 9.75183 | 0.504791 | 0.0789154 |
| ACC 6h | 78.4253 | 7.89771 | 0.799814 | 0.145016 |
| BABA 6h | 114.861 | 7.91809 | 1.20806 | 0.101133 |
| Chitin 6h | 119.764 | 8.11922 | 1.32412 | 0.0896316 |
| Epi 6h | 133.42 | 8.86296 | 1.39166 | 0.0919385 |
| SA 6h | 128.544 | 8.48527 | 1.52886 | 0.166094 |
| Me-JA 6h | 93.7221 | 25.5455 | 1.03865 | 0.220048 |
Source Transcript PGSC0003DMT400061439 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G07710.1 | +2 | 0.0 | 592 | 335/540 (62%) | Ankyrin repeat family protein | chr1:2386275-2387986 REVERSE LENGTH=543 |
