Probe CUST_13173_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_13173_PI426222305 | JHI_St_60k_v1 | DMT400007477 | TTTTTGACGGCATCTTTCCCAAGAAAATGAGAAGGCCAAAGAAGAGAGCTCAGAGACGAC |
All Microarray Probes Designed to Gene DMG400002885
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_13121_PI426222305 | JHI_St_60k_v1 | DMT400007478 | GAAAAGTGTCAATGTCCATTTTTTGGTCATTTTTTACTTCATTATCAAGGGATTGGGCTG |
| CUST_13173_PI426222305 | JHI_St_60k_v1 | DMT400007477 | TTTTTGACGGCATCTTTCCCAAGAAAATGAGAAGGCCAAAGAAGAGAGCTCAGAGACGAC |
Microarray Signals from CUST_13173_PI426222305
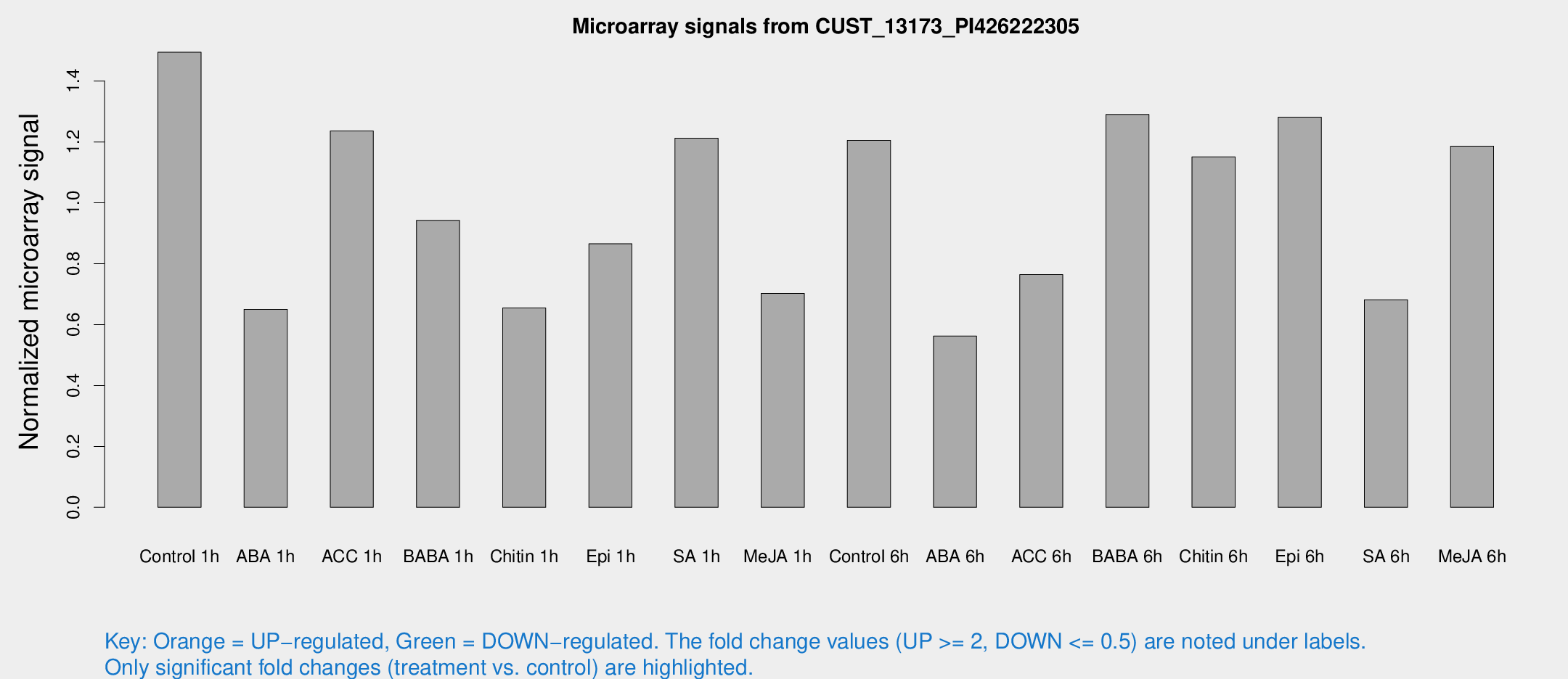
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 136.871 | 48.9232 | 1.49452 | 0.430887 |
| ABA 1h | 47.7361 | 5.8553 | 0.650258 | 0.132981 |
| ACC 1h | 111.687 | 30.7661 | 1.23582 | 0.283301 |
| BABA 1h | 80.9399 | 21.3654 | 0.942358 | 0.204567 |
| Chitin 1h | 58.0777 | 23.4675 | 0.654509 | 0.272696 |
| Epi 1h | 64.6207 | 15.3231 | 0.865461 | 0.187116 |
| SA 1h | 111.741 | 36.1331 | 1.21202 | 0.308319 |
| Me-JA 1h | 49.7498 | 12.8747 | 0.702324 | 0.129145 |
| Control 6h | 103.758 | 21.2295 | 1.20491 | 0.174295 |
| ABA 6h | 55.2565 | 17.3332 | 0.562162 | 0.195408 |
| ACC 6h | 73.3255 | 11.4606 | 0.764254 | 0.154896 |
| BABA 6h | 118.84 | 10.6677 | 1.28997 | 0.121282 |
| Chitin 6h | 104.271 | 20.5096 | 1.15084 | 0.20944 |
| Epi 6h | 121.375 | 21.2004 | 1.28118 | 0.201536 |
| SA 6h | 75.0732 | 31.8067 | 0.68168 | 0.435072 |
| Me-JA 6h | 108.436 | 36.8894 | 1.18616 | 0.324922 |
Source Transcript PGSC0003DMT400007477 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G45616.1 | +1 | 2e-142 | 452 | 340/958 (35%) | receptor like protein 6 | chr1:17183550-17186534 REVERSE LENGTH=994 |
