Probe CUST_13121_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_13121_PI426222305 | JHI_St_60k_v1 | DMT400007478 | GAAAAGTGTCAATGTCCATTTTTTGGTCATTTTTTACTTCATTATCAAGGGATTGGGCTG |
All Microarray Probes Designed to Gene DMG400002885
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_13121_PI426222305 | JHI_St_60k_v1 | DMT400007478 | GAAAAGTGTCAATGTCCATTTTTTGGTCATTTTTTACTTCATTATCAAGGGATTGGGCTG |
| CUST_13173_PI426222305 | JHI_St_60k_v1 | DMT400007477 | TTTTTGACGGCATCTTTCCCAAGAAAATGAGAAGGCCAAAGAAGAGAGCTCAGAGACGAC |
Microarray Signals from CUST_13121_PI426222305
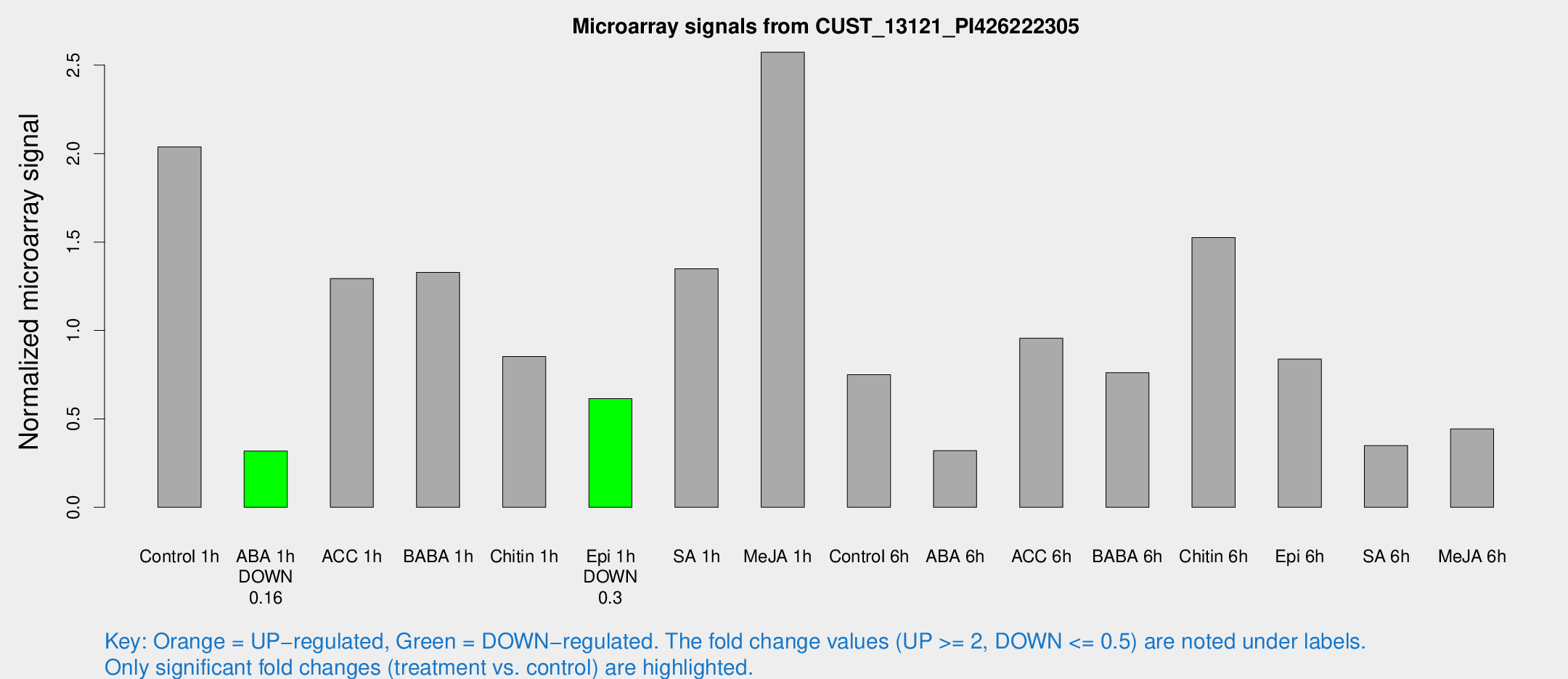
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 50.389 | 12.2487 | 2.03812 | 0.3465 |
| ABA 1h | 6.64139 | 3.47057 | 0.31799 | 0.169208 |
| ACC 1h | 38.9892 | 14.2188 | 1.2932 | 0.697533 |
| BABA 1h | 31.1422 | 6.37703 | 1.32874 | 0.204898 |
| Chitin 1h | 21.0722 | 8.84075 | 0.852928 | 0.370612 |
| Epi 1h | 12.4974 | 3.544 | 0.615046 | 0.175298 |
| SA 1h | 33.8397 | 7.11761 | 1.34896 | 0.262399 |
| Me-JA 1h | 53.1545 | 15.2006 | 2.57314 | 0.50553 |
| Control 6h | 21.0994 | 7.26041 | 0.749749 | 0.317581 |
| ABA 6h | 8.19458 | 3.88602 | 0.320505 | 0.161308 |
| ACC 6h | 27.0521 | 6.08687 | 0.956245 | 0.167208 |
| BABA 6h | 23.7507 | 8.05377 | 0.760954 | 0.457673 |
| Chitin 6h | 60.7426 | 39.4566 | 1.52503 | 1.63257 |
| Epi 6h | 25.4921 | 7.90839 | 0.837815 | 0.477904 |
| SA 6h | 8.26949 | 4.0395 | 0.3486 | 0.178144 |
| Me-JA 6h | 10.9213 | 3.73952 | 0.44333 | 0.174516 |
Source Transcript PGSC0003DMT400007478 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G45616.1 | +1 | 9e-86 | 300 | 202/504 (40%) | receptor like protein 6 | chr1:17183550-17186534 REVERSE LENGTH=994 |
