Probe CUST_12511_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_12511_PI426222305 | JHI_St_60k_v1 | DMT400063588 | ACAGGGAGTTTCTTCTAGCTAGTTACATGCATGGTGAGTACTCTGTAGAGTGAGACTAAT |
All Microarray Probes Designed to Gene DMG400024720
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_12511_PI426222305 | JHI_St_60k_v1 | DMT400063588 | ACAGGGAGTTTCTTCTAGCTAGTTACATGCATGGTGAGTACTCTGTAGAGTGAGACTAAT |
| CUST_12573_PI426222305 | JHI_St_60k_v1 | DMT400063589 | AAGGTCCAAGTTCGTCATGTATTCTGCTTTGTATTACGAGTTTCCCTTCTAATGTTATTT |
Microarray Signals from CUST_12511_PI426222305
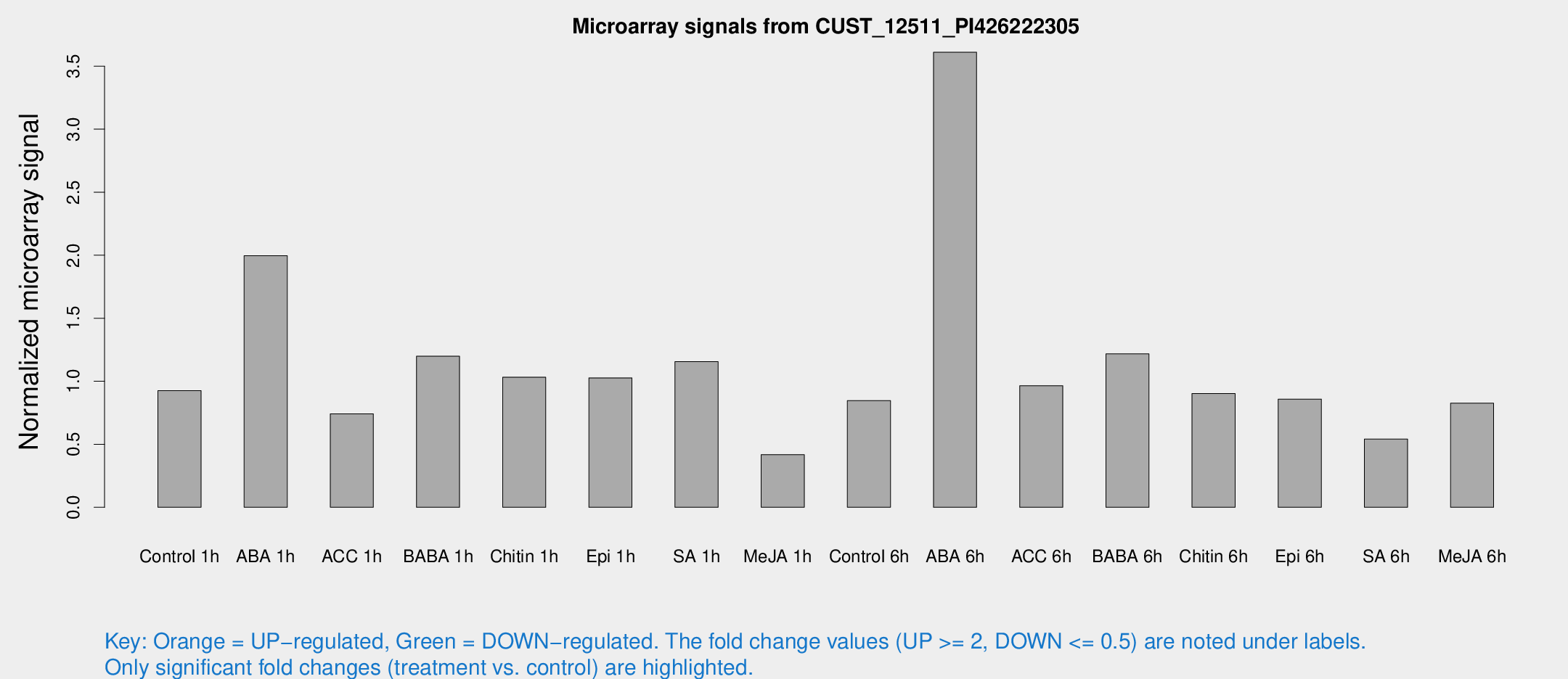
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 284.388 | 39.0615 | 0.925695 | 0.0867479 |
| ABA 1h | 559.364 | 119.072 | 1.99633 | 0.314866 |
| ACC 1h | 251.078 | 64.6764 | 0.741531 | 0.184359 |
| BABA 1h | 380.785 | 100.343 | 1.19843 | 0.260105 |
| Chitin 1h | 281.385 | 18.5095 | 1.03226 | 0.10031 |
| Epi 1h | 284.004 | 60.8463 | 1.02704 | 0.251436 |
| SA 1h | 368.252 | 59.748 | 1.15608 | 0.176716 |
| Me-JA 1h | 106.347 | 19.7409 | 0.418137 | 0.0491108 |
| Control 6h | 278.124 | 76.3438 | 0.846873 | 0.192915 |
| ABA 6h | 1214 | 250.288 | 3.61071 | 0.618662 |
| ACC 6h | 334.416 | 19.7349 | 0.965081 | 0.10428 |
| BABA 6h | 437.056 | 108.391 | 1.21745 | 0.260701 |
| Chitin 6h | 292.721 | 27.7374 | 0.902632 | 0.0853881 |
| Epi 6h | 310.025 | 67.7635 | 0.858718 | 0.162452 |
| SA 6h | 218.554 | 111.369 | 0.541358 | 0.301402 |
| Me-JA 6h | 285.632 | 89.1778 | 0.825965 | 0.302248 |
Source Transcript PGSC0003DMT400063588 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G56220.3 | +1 | 5e-29 | 113 | 61/98 (62%) | Dormancy/auxin associated family protein | chr1:21043414-21044469 FORWARD LENGTH=140 |
