Probe CUST_12573_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_12573_PI426222305 | JHI_St_60k_v1 | DMT400063589 | AAGGTCCAAGTTCGTCATGTATTCTGCTTTGTATTACGAGTTTCCCTTCTAATGTTATTT |
All Microarray Probes Designed to Gene DMG400024720
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_12511_PI426222305 | JHI_St_60k_v1 | DMT400063588 | ACAGGGAGTTTCTTCTAGCTAGTTACATGCATGGTGAGTACTCTGTAGAGTGAGACTAAT |
| CUST_12573_PI426222305 | JHI_St_60k_v1 | DMT400063589 | AAGGTCCAAGTTCGTCATGTATTCTGCTTTGTATTACGAGTTTCCCTTCTAATGTTATTT |
Microarray Signals from CUST_12573_PI426222305
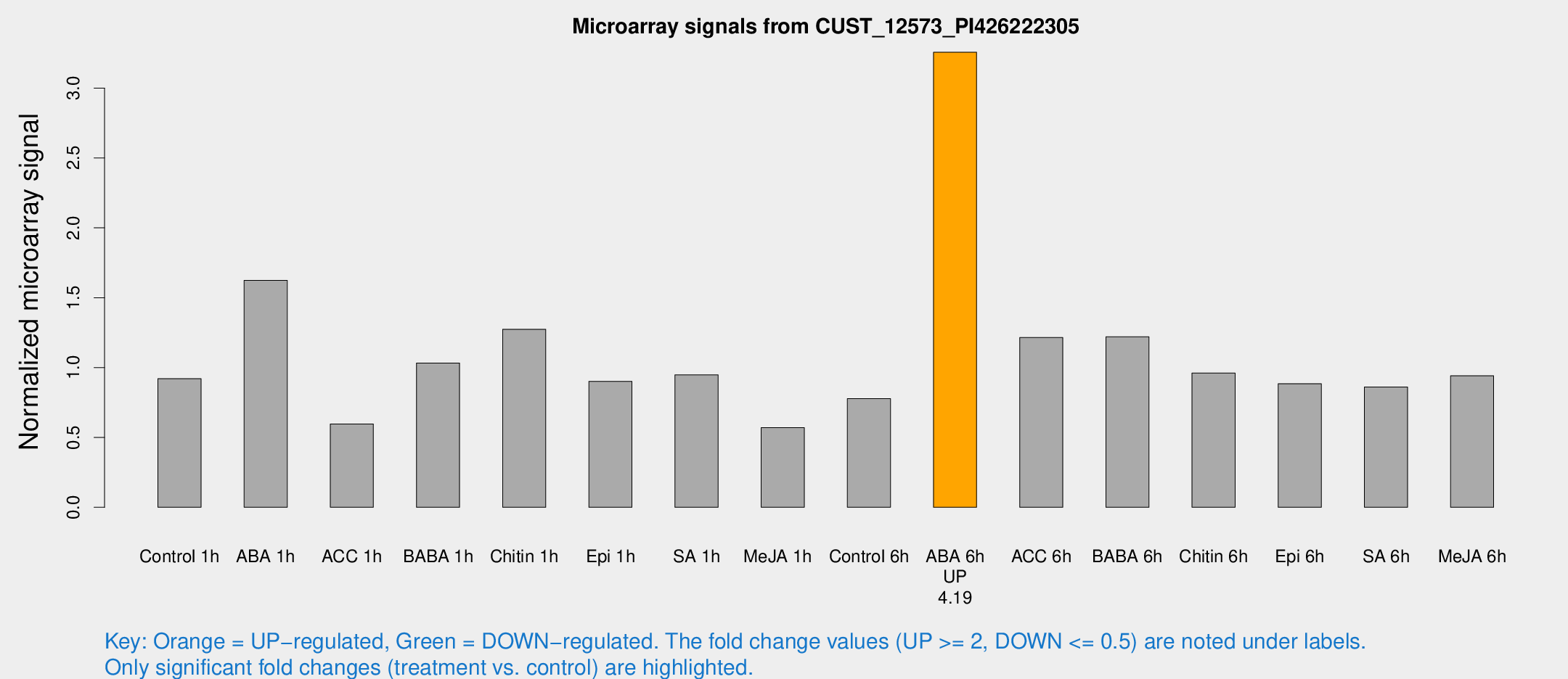
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 23314.6 | 1347.33 | 0.920165 | 0.0531259 |
| ABA 1h | 38528.4 | 8790.21 | 1.6242 | 0.296721 |
| ACC 1h | 16121.1 | 2943.67 | 0.596326 | 0.0755936 |
| BABA 1h | 27792 | 7628.41 | 1.03274 | 0.244873 |
| Chitin 1h | 29232.8 | 2646.88 | 1.27403 | 0.0916662 |
| Epi 1h | 20459.9 | 3534.32 | 0.901755 | 0.163605 |
| SA 1h | 25757.5 | 5555.71 | 0.947433 | 0.170211 |
| Me-JA 1h | 11982 | 1594.32 | 0.57023 | 0.0445774 |
| Control 6h | 21025.9 | 5445.71 | 0.777558 | 0.161451 |
| ABA 6h | 89085.3 | 11233.4 | 3.25673 | 0.239437 |
| ACC 6h | 35349.4 | 2044.18 | 1.21536 | 0.125906 |
| BABA 6h | 35768.2 | 6812.14 | 1.21977 | 0.182031 |
| Chitin 6h | 25956.2 | 1502.4 | 0.960859 | 0.0554755 |
| Epi 6h | 26292.1 | 5385.58 | 0.884664 | 0.113614 |
| SA 6h | 22335.4 | 3867.15 | 0.861434 | 0.0657136 |
| Me-JA 6h | 24770.8 | 4781.96 | 0.940741 | 0.128277 |
Source Transcript PGSC0003DMT400063589 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G56220.3 | +1 | 1e-28 | 108 | 58/92 (63%) | Dormancy/auxin associated family protein | chr1:21043414-21044469 FORWARD LENGTH=140 |
