Probe CUST_11856_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_11856_PI426222305 | JHI_St_60k_v1 | DMT400046696 | CTGAGTTTGATGACTCATCATGTGTCATGCGTGACATTGTTAATTACCTTCTTGTTTTTA |
All Microarray Probes Designed to Gene DMG400018129
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_11856_PI426222305 | JHI_St_60k_v1 | DMT400046696 | CTGAGTTTGATGACTCATCATGTGTCATGCGTGACATTGTTAATTACCTTCTTGTTTTTA |
| CUST_11783_PI426222305 | JHI_St_60k_v1 | DMT400046695 | CTAGTCGGATTCTACTTCGTTGAGAAGAGAAAGGCGGTCAAAATTTCCCAGCGAAAATGA |
Microarray Signals from CUST_11856_PI426222305
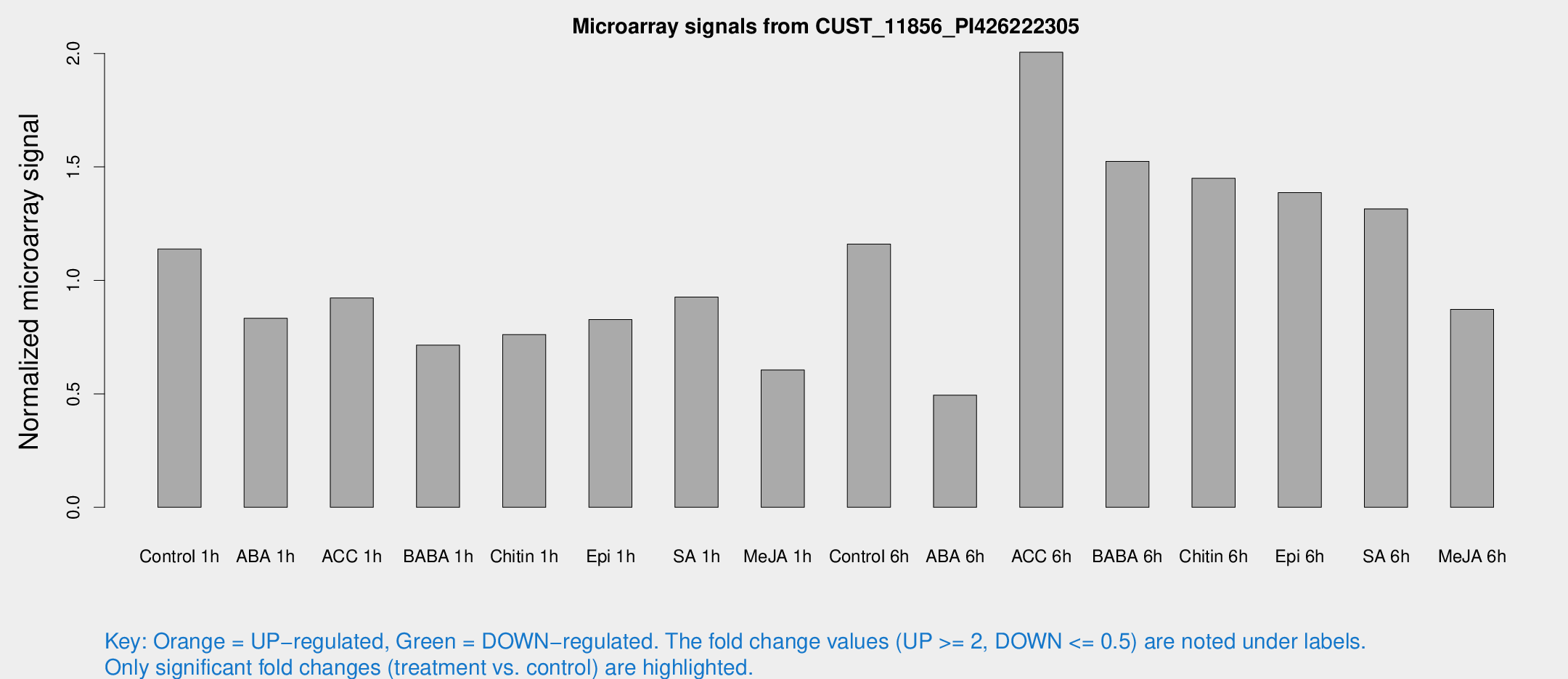
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 209.431 | 40.9253 | 1.13819 | 0.163872 |
| ABA 1h | 131.253 | 8.31994 | 0.833248 | 0.0934775 |
| ACC 1h | 183.518 | 47.4544 | 0.922434 | 0.227809 |
| BABA 1h | 139.855 | 51.3777 | 0.714672 | 0.217105 |
| Chitin 1h | 121.649 | 7.85812 | 0.761267 | 0.0683254 |
| Epi 1h | 127.596 | 8.13357 | 0.82754 | 0.0685413 |
| SA 1h | 169.793 | 11.4361 | 0.926459 | 0.0569297 |
| Me-JA 1h | 92.8346 | 21.6152 | 0.605158 | 0.0945461 |
| Control 6h | 244.267 | 80.7355 | 1.15957 | 0.459271 |
| ABA 6h | 96.3655 | 16.0001 | 0.494375 | 0.0701227 |
| ACC 6h | 413.631 | 44.3753 | 2.00534 | 0.270206 |
| BABA 6h | 305.621 | 28.2366 | 1.52448 | 0.0903938 |
| Chitin 6h | 276.281 | 25.6708 | 1.44958 | 0.101601 |
| Epi 6h | 284.021 | 41.0359 | 1.38676 | 0.148135 |
| SA 6h | 232.623 | 20.1773 | 1.31494 | 0.123817 |
| Me-JA 6h | 157.637 | 23.1618 | 0.872393 | 0.106089 |
Source Transcript PGSC0003DMT400046696 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G50200.3 | +1 | 1e-48 | 164 | 89/184 (48%) | nitrate transmembrane transporters | chr5:20436308-20437535 FORWARD LENGTH=210 |
