Probe CUST_11783_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_11783_PI426222305 | JHI_St_60k_v1 | DMT400046695 | CTAGTCGGATTCTACTTCGTTGAGAAGAGAAAGGCGGTCAAAATTTCCCAGCGAAAATGA |
All Microarray Probes Designed to Gene DMG400018129
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_11856_PI426222305 | JHI_St_60k_v1 | DMT400046696 | CTGAGTTTGATGACTCATCATGTGTCATGCGTGACATTGTTAATTACCTTCTTGTTTTTA |
| CUST_11783_PI426222305 | JHI_St_60k_v1 | DMT400046695 | CTAGTCGGATTCTACTTCGTTGAGAAGAGAAAGGCGGTCAAAATTTCCCAGCGAAAATGA |
Microarray Signals from CUST_11783_PI426222305
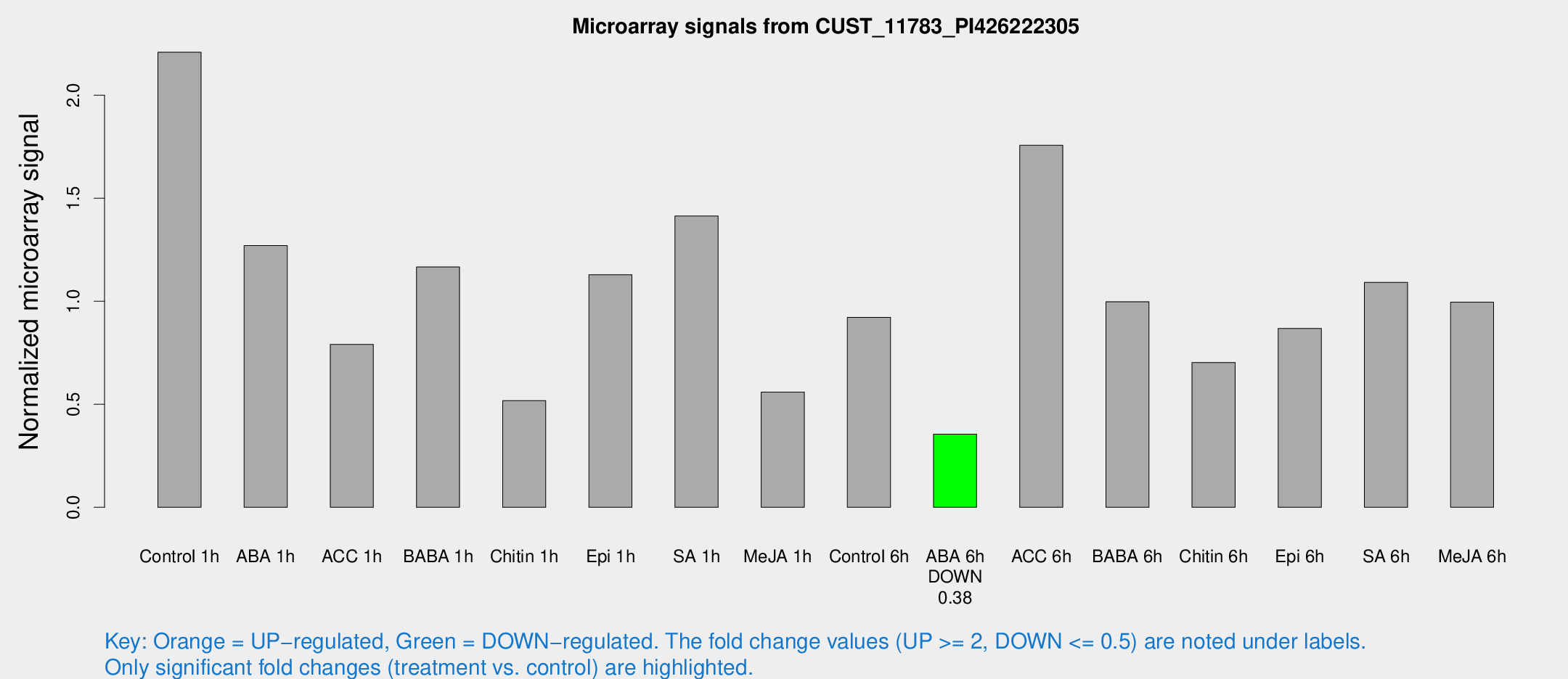
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 3996.06 | 456.473 | 2.20877 | 0.209137 |
| ABA 1h | 2059.59 | 337.446 | 1.27083 | 0.202916 |
| ACC 1h | 2204.99 | 1021.11 | 0.7904 | 0.801183 |
| BABA 1h | 2150.71 | 510.442 | 1.16628 | 0.2041 |
| Chitin 1h | 901.096 | 222.571 | 0.518075 | 0.171998 |
| Epi 1h | 1819.08 | 364.697 | 1.12916 | 0.234713 |
| SA 1h | 2598.11 | 150.286 | 1.41383 | 0.0816482 |
| Me-JA 1h | 873.269 | 232.753 | 0.559526 | 0.12236 |
| Control 6h | 1704.17 | 316.526 | 0.921884 | 0.137487 |
| ABA 6h | 684.888 | 85.5356 | 0.354799 | 0.037707 |
| ACC 6h | 3634.63 | 361.572 | 1.75754 | 0.377856 |
| BABA 6h | 2071.55 | 419.436 | 0.997856 | 0.160685 |
| Chitin 6h | 1338.18 | 77.4688 | 0.703104 | 0.0406576 |
| Epi 6h | 1778.47 | 230.349 | 0.868181 | 0.0501686 |
| SA 6h | 1968.04 | 266.884 | 1.09181 | 0.0630799 |
| Me-JA 6h | 1781.12 | 139.428 | 0.995669 | 0.161654 |
Source Transcript PGSC0003DMT400046695 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G50200.3 | +1 | 2e-53 | 177 | 88/179 (49%) | nitrate transmembrane transporters | chr5:20436308-20437535 FORWARD LENGTH=210 |
