Probe CUST_11640_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_11640_PI426222305 | JHI_St_60k_v1 | DMT400008122 | CGGAGGGAGTATCATTCAATGAACTGCTTGATATCTGACACTTTGTTGTTACATTTTATT |
All Microarray Probes Designed to Gene DMG400003133
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_11640_PI426222305 | JHI_St_60k_v1 | DMT400008122 | CGGAGGGAGTATCATTCAATGAACTGCTTGATATCTGACACTTTGTTGTTACATTTTATT |
| CUST_11630_PI426222305 | JHI_St_60k_v1 | DMT400008123 | CAAAAATAATTACTTCTTGTGGCTTTGCGGCCCTTATCGTCTTGTTTGTGGAGGAGGGGA |
Microarray Signals from CUST_11640_PI426222305
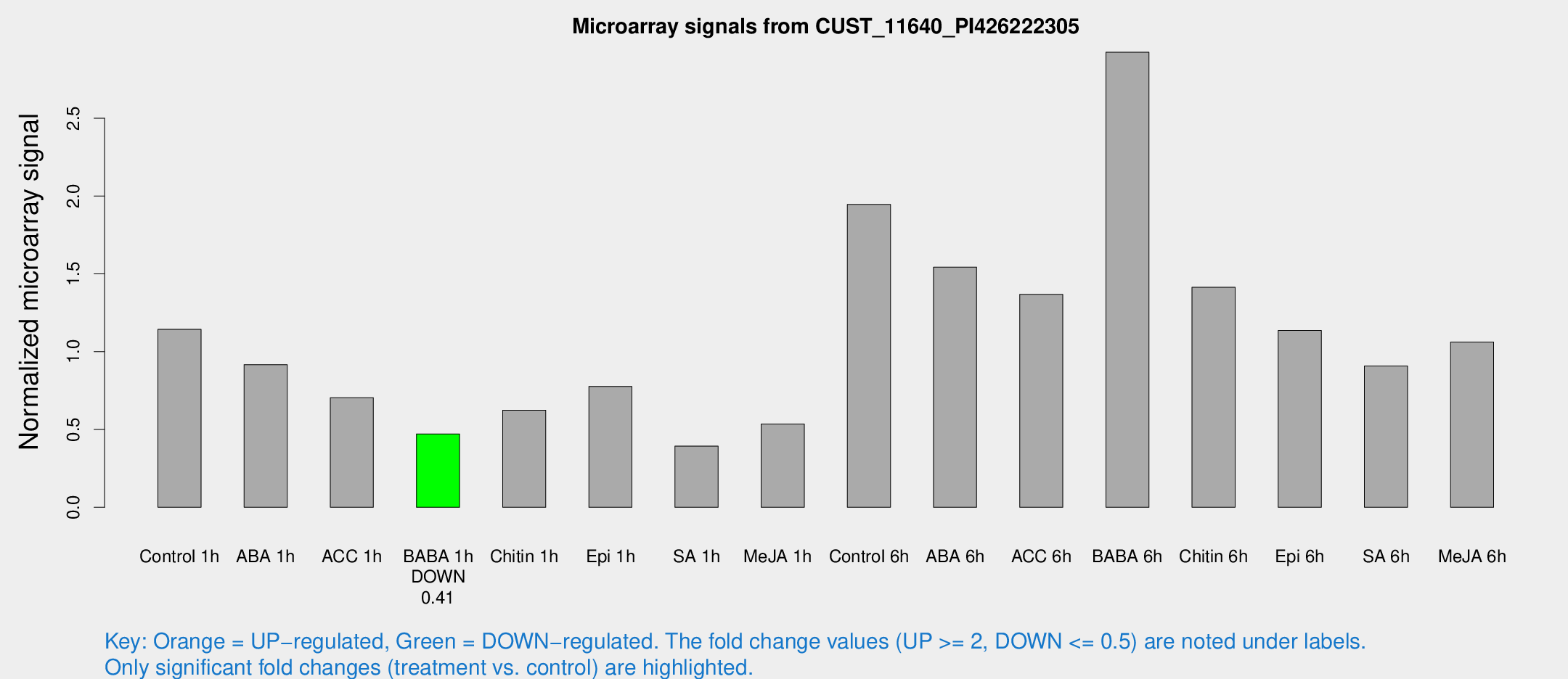
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 28.0191 | 3.73135 | 1.14374 | 0.152623 |
| ABA 1h | 20.2001 | 3.46049 | 0.916874 | 0.165281 |
| ACC 1h | 19.9194 | 6.11404 | 0.70462 | 0.217514 |
| BABA 1h | 11.1486 | 3.62901 | 0.471545 | 0.154152 |
| Chitin 1h | 14.3819 | 3.51447 | 0.624247 | 0.198015 |
| Epi 1h | 16.6789 | 3.51129 | 0.776723 | 0.16863 |
| SA 1h | 11.7092 | 4.97808 | 0.393437 | 0.213157 |
| Me-JA 1h | 11.7085 | 3.60007 | 0.535356 | 0.205304 |
| Control 6h | 54.5316 | 19.1187 | 1.94626 | 0.62708 |
| ABA 6h | 44.5608 | 14.7488 | 1.54362 | 0.424501 |
| ACC 6h | 39.8743 | 7.47049 | 1.36834 | 0.298092 |
| BABA 6h | 132.193 | 82.0498 | 2.92435 | 3.24231 |
| Chitin 6h | 46.2123 | 17.2453 | 1.41391 | 0.954588 |
| Epi 6h | 44.5041 | 18.9457 | 1.13607 | 1.3639 |
| SA 6h | 22.8205 | 4.20291 | 0.90806 | 0.292181 |
| Me-JA 6h | 37.2792 | 15.7917 | 1.06223 | 0.923967 |
Source Transcript PGSC0003DMT400008122 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G53170.1 | +2 | 0.0 | 566 | 263/413 (64%) | Tetratricopeptide repeat (TPR)-like superfamily protein | chr3:19704600-19706417 REVERSE LENGTH=499 |
