Probe CUST_11630_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_11630_PI426222305 | JHI_St_60k_v1 | DMT400008123 | CAAAAATAATTACTTCTTGTGGCTTTGCGGCCCTTATCGTCTTGTTTGTGGAGGAGGGGA |
All Microarray Probes Designed to Gene DMG400003133
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_11640_PI426222305 | JHI_St_60k_v1 | DMT400008122 | CGGAGGGAGTATCATTCAATGAACTGCTTGATATCTGACACTTTGTTGTTACATTTTATT |
| CUST_11630_PI426222305 | JHI_St_60k_v1 | DMT400008123 | CAAAAATAATTACTTCTTGTGGCTTTGCGGCCCTTATCGTCTTGTTTGTGGAGGAGGGGA |
Microarray Signals from CUST_11630_PI426222305
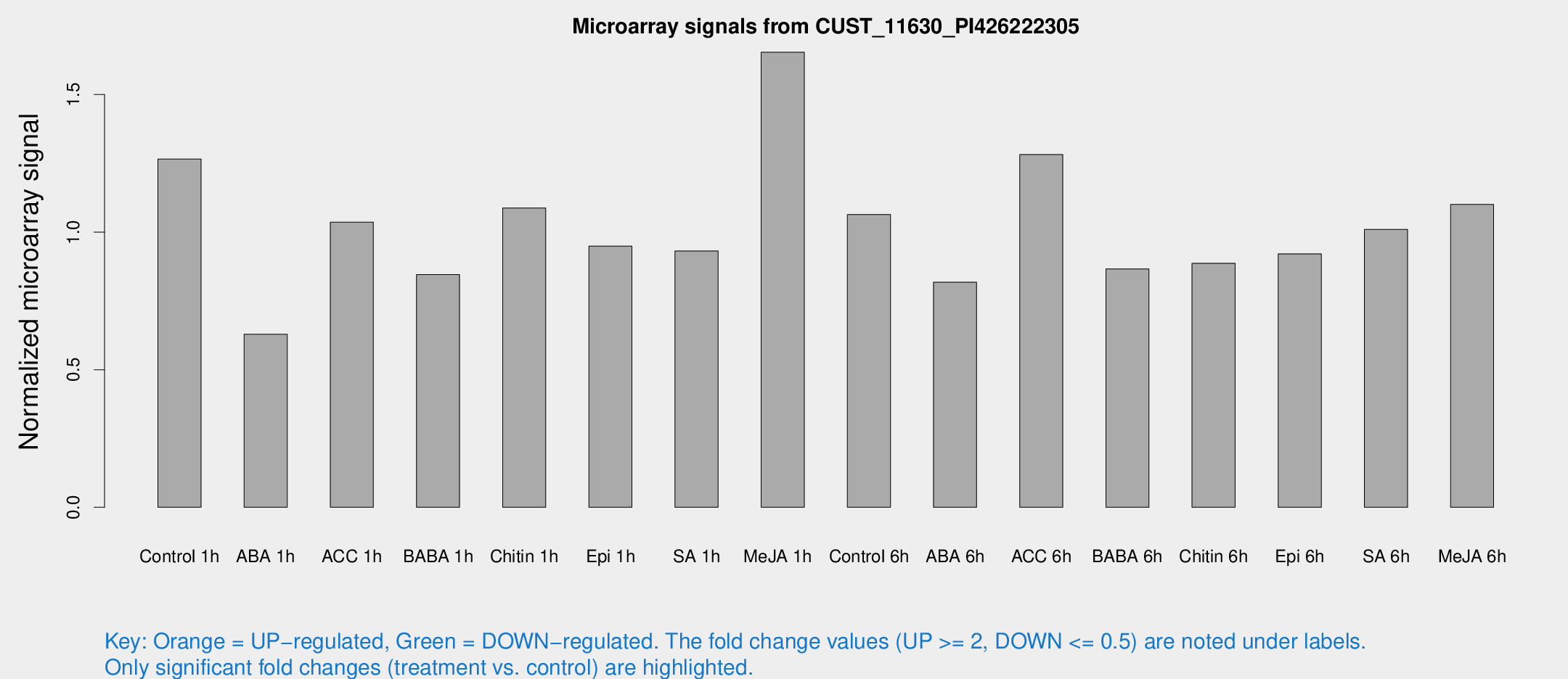
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 21.9111 | 3.49772 | 1.26538 | 0.277392 |
| ABA 1h | 9.789 | 3.00477 | 0.628874 | 0.21427 |
| ACC 1h | 18.1001 | 3.84587 | 1.03598 | 0.221414 |
| BABA 1h | 14.3739 | 3.38561 | 0.845857 | 0.222786 |
| Chitin 1h | 16.8546 | 3.21457 | 1.08787 | 0.212468 |
| Epi 1h | 14.0116 | 3.11379 | 0.949506 | 0.219577 |
| SA 1h | 17.4537 | 4.76324 | 0.931746 | 0.322401 |
| Me-JA 1h | 23.6691 | 4.7645 | 1.65379 | 0.259152 |
| Control 6h | 18.8841 | 3.89584 | 1.06414 | 0.337576 |
| ABA 6h | 15.205 | 3.38788 | 0.817897 | 0.197283 |
| ACC 6h | 25.2278 | 4.0403 | 1.28191 | 0.201438 |
| BABA 6h | 17.7823 | 5.39845 | 0.866382 | 0.297815 |
| Chitin 6h | 16.6473 | 3.68696 | 0.886967 | 0.216263 |
| Epi 6h | 17.5719 | 3.81855 | 0.921203 | 0.201977 |
| SA 6h | 16.8762 | 3.50451 | 1.00992 | 0.210539 |
| Me-JA 6h | 18.6521 | 3.31753 | 1.10053 | 0.199341 |
Source Transcript PGSC0003DMT400008123 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G53170.1 | +2 | 0.0 | 566 | 263/413 (64%) | Tetratricopeptide repeat (TPR)-like superfamily protein | chr3:19704600-19706417 REVERSE LENGTH=499 |
