Probe CUST_11209_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_11209_PI426222305 | JHI_St_60k_v1 | DMT400007377 | GGTCAAACGTGTCTTTCAGAGTTCAACTTTGAAAAAAGTGAAAAGATTTCACAACCAAAC |
All Microarray Probes Designed to Gene DMG400002835
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_11209_PI426222305 | JHI_St_60k_v1 | DMT400007377 | GGTCAAACGTGTCTTTCAGAGTTCAACTTTGAAAAAAGTGAAAAGATTTCACAACCAAAC |
| CUST_11051_PI426222305 | JHI_St_60k_v1 | DMT400007376 | GAGAATGAGAGCTTGAGAAATGAGGTACATTTACTCAAAGATAAGTTGATCAACAGCGTA |
Microarray Signals from CUST_11209_PI426222305
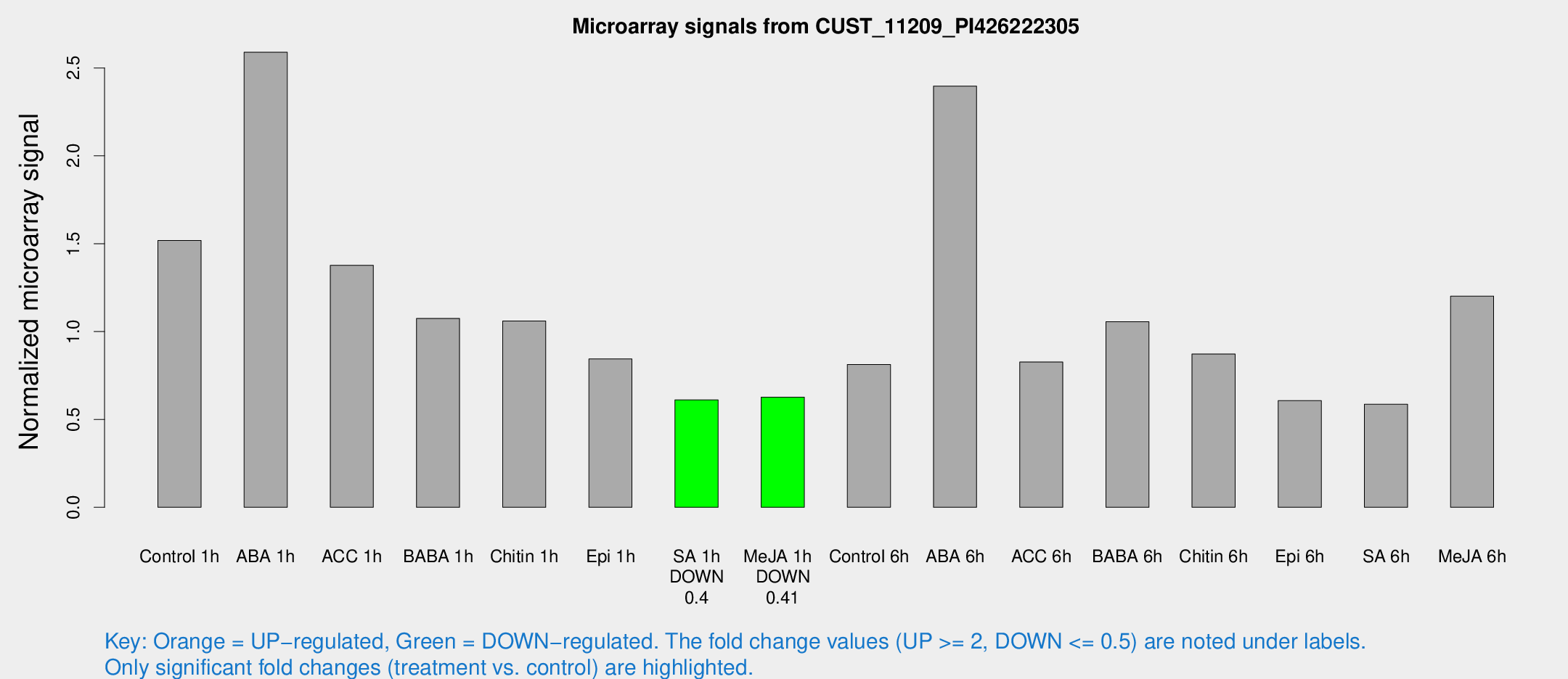
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 121.081 | 14.4685 | 1.51816 | 0.102817 |
| ABA 1h | 183.409 | 24.2059 | 2.58937 | 0.228184 |
| ACC 1h | 115.223 | 20.5678 | 1.37703 | 0.172432 |
| BABA 1h | 83.0095 | 11.8991 | 1.07468 | 0.183166 |
| Chitin 1h | 78.1359 | 16.4032 | 1.05973 | 0.185562 |
| Epi 1h | 59.6483 | 10.8066 | 0.844003 | 0.164705 |
| SA 1h | 50.8431 | 8.29534 | 0.611282 | 0.0767452 |
| Me-JA 1h | 42.0049 | 8.35546 | 0.62717 | 0.0822748 |
| Control 6h | 72.0834 | 25.0529 | 0.812899 | 0.24631 |
| ABA 6h | 208.863 | 40.0355 | 2.39674 | 0.40214 |
| ACC 6h | 79.8291 | 21.1103 | 0.826744 | 0.113898 |
| BABA 6h | 94.4673 | 11.94 | 1.05599 | 0.107946 |
| Chitin 6h | 75.8612 | 14.7244 | 0.87221 | 0.21222 |
| Epi 6h | 56.3259 | 12.6628 | 0.606986 | 0.125954 |
| SA 6h | 45.5761 | 4.49565 | 0.586417 | 0.0640435 |
| Me-JA 6h | 97.9879 | 18.9817 | 1.2013 | 0.15356 |
Source Transcript PGSC0003DMT400007377 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G01470.1 | +3 | 2e-33 | 131 | 62/91 (68%) | homeobox 1 | chr3:182648-184034 REVERSE LENGTH=272 |
