Probe CUST_11051_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_11051_PI426222305 | JHI_St_60k_v1 | DMT400007376 | GAGAATGAGAGCTTGAGAAATGAGGTACATTTACTCAAAGATAAGTTGATCAACAGCGTA |
All Microarray Probes Designed to Gene DMG400002835
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_11209_PI426222305 | JHI_St_60k_v1 | DMT400007377 | GGTCAAACGTGTCTTTCAGAGTTCAACTTTGAAAAAAGTGAAAAGATTTCACAACCAAAC |
| CUST_11051_PI426222305 | JHI_St_60k_v1 | DMT400007376 | GAGAATGAGAGCTTGAGAAATGAGGTACATTTACTCAAAGATAAGTTGATCAACAGCGTA |
Microarray Signals from CUST_11051_PI426222305
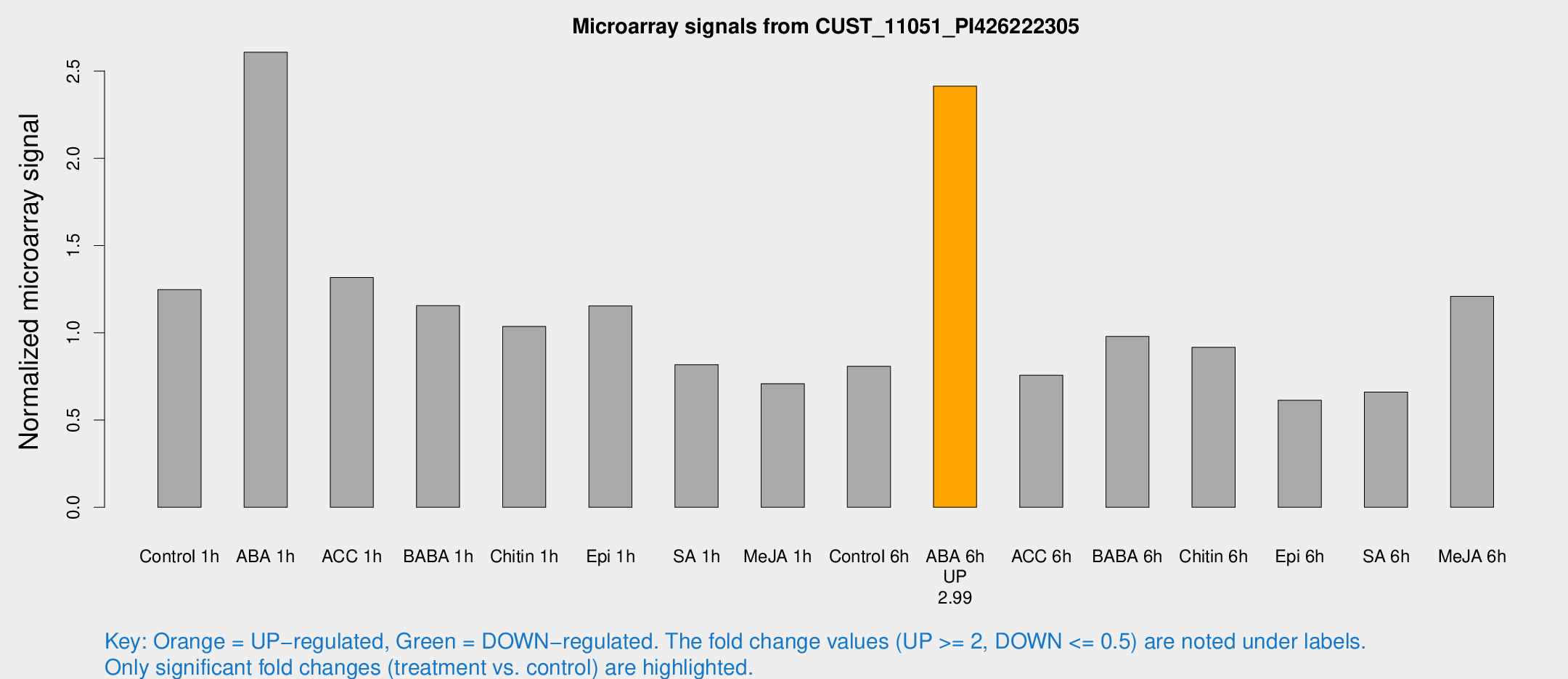
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 711.748 | 59.2932 | 1.24777 | 0.0722307 |
| ABA 1h | 1385.45 | 322.308 | 2.60811 | 0.470377 |
| ACC 1h | 807.23 | 170.564 | 1.31675 | 0.220552 |
| BABA 1h | 666.139 | 139.976 | 1.15576 | 0.160413 |
| Chitin 1h | 563.575 | 150.398 | 1.03618 | 0.206665 |
| Epi 1h | 588.857 | 110.117 | 1.15405 | 0.218388 |
| SA 1h | 496.711 | 103.616 | 0.817389 | 0.120242 |
| Me-JA 1h | 342.085 | 68.891 | 0.708541 | 0.0879624 |
| Control 6h | 496.68 | 131.653 | 0.807661 | 0.177792 |
| ABA 6h | 1485.31 | 209.376 | 2.41404 | 0.228096 |
| ACC 6h | 506.133 | 86.5181 | 0.757088 | 0.0440418 |
| BABA 6h | 624.686 | 46.7353 | 0.979109 | 0.0567786 |
| Chitin 6h | 556.882 | 44.209 | 0.917239 | 0.0974159 |
| Epi 6h | 392.328 | 22.9278 | 0.613746 | 0.0591384 |
| SA 6h | 390.217 | 83.5933 | 0.660793 | 0.0853377 |
| Me-JA 6h | 702.061 | 120.235 | 1.2095 | 0.106943 |
Source Transcript PGSC0003DMT400007376 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G01470.1 | +1 | 2e-40 | 149 | 97/214 (45%) | homeobox 1 | chr3:182648-184034 REVERSE LENGTH=272 |
