Probe CUST_10880_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10880_PI426222305 | JHI_St_60k_v1 | DMT400031782 | GCTCGTAATGATGAAGTTGGTCTTGAGATTCCGAAAACTTCATCATCAACATTGTTTAAT |
All Microarray Probes Designed to Gene DMG400012193
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10792_PI426222305 | JHI_St_60k_v1 | DMT400031783 | CTATGGAGGGAATAGACTAGCTATAGCTTATCGAGCACAAATTGTTTTTCTGTCTTTTTT |
| CUST_10880_PI426222305 | JHI_St_60k_v1 | DMT400031782 | GCTCGTAATGATGAAGTTGGTCTTGAGATTCCGAAAACTTCATCATCAACATTGTTTAAT |
Microarray Signals from CUST_10880_PI426222305
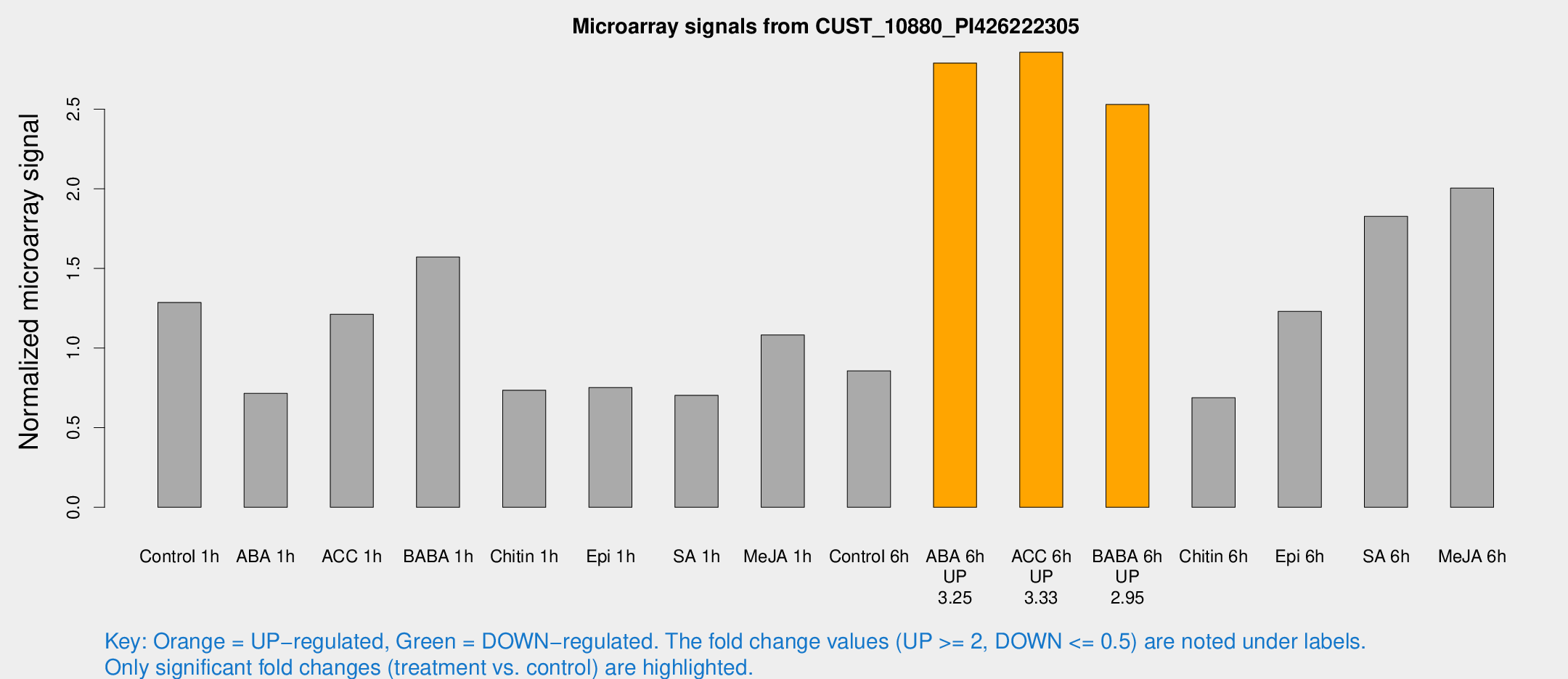
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 11.3647 | 3.10915 | 1.28631 | 0.464334 |
| ABA 1h | 4.93128 | 2.8611 | 0.715407 | 0.40359 |
| ACC 1h | 13.9139 | 8.20337 | 1.21196 | 0.811737 |
| BABA 1h | 13.9471 | 4.32843 | 1.57125 | 0.532587 |
| Chitin 1h | 5.14908 | 2.99513 | 0.735129 | 0.413581 |
| Epi 1h | 5.1484 | 2.98674 | 0.752362 | 0.429824 |
| SA 1h | 5.85867 | 2.98917 | 0.702909 | 0.368738 |
| Me-JA 1h | 7.77466 | 3.07784 | 1.08212 | 0.503961 |
| Control 6h | 7.33943 | 3.10393 | 0.857144 | 0.414735 |
| ABA 6h | 24.4872 | 4.53001 | 2.78924 | 0.528918 |
| ACC 6h | 27.0216 | 4.54476 | 2.85709 | 0.952356 |
| BABA 6h | 23.1693 | 3.72337 | 2.52896 | 0.419759 |
| Chitin 6h | 5.86138 | 3.40361 | 0.687958 | 0.399404 |
| Epi 6h | 11.1702 | 3.6113 | 1.23023 | 0.404632 |
| SA 6h | 17.6139 | 6.83412 | 1.8262 | 0.733465 |
| Me-JA 6h | 16.3572 | 3.22568 | 2.00427 | 0.414696 |
Source Transcript PGSC0003DMT400031782 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G63820.1 | +1 | 4e-47 | 161 | 88/156 (56%) | CCT motif family protein | chr1:23682529-23684050 REVERSE LENGTH=293 |
