Probe CUST_10792_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10792_PI426222305 | JHI_St_60k_v1 | DMT400031783 | CTATGGAGGGAATAGACTAGCTATAGCTTATCGAGCACAAATTGTTTTTCTGTCTTTTTT |
All Microarray Probes Designed to Gene DMG400012193
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10792_PI426222305 | JHI_St_60k_v1 | DMT400031783 | CTATGGAGGGAATAGACTAGCTATAGCTTATCGAGCACAAATTGTTTTTCTGTCTTTTTT |
| CUST_10880_PI426222305 | JHI_St_60k_v1 | DMT400031782 | GCTCGTAATGATGAAGTTGGTCTTGAGATTCCGAAAACTTCATCATCAACATTGTTTAAT |
Microarray Signals from CUST_10792_PI426222305
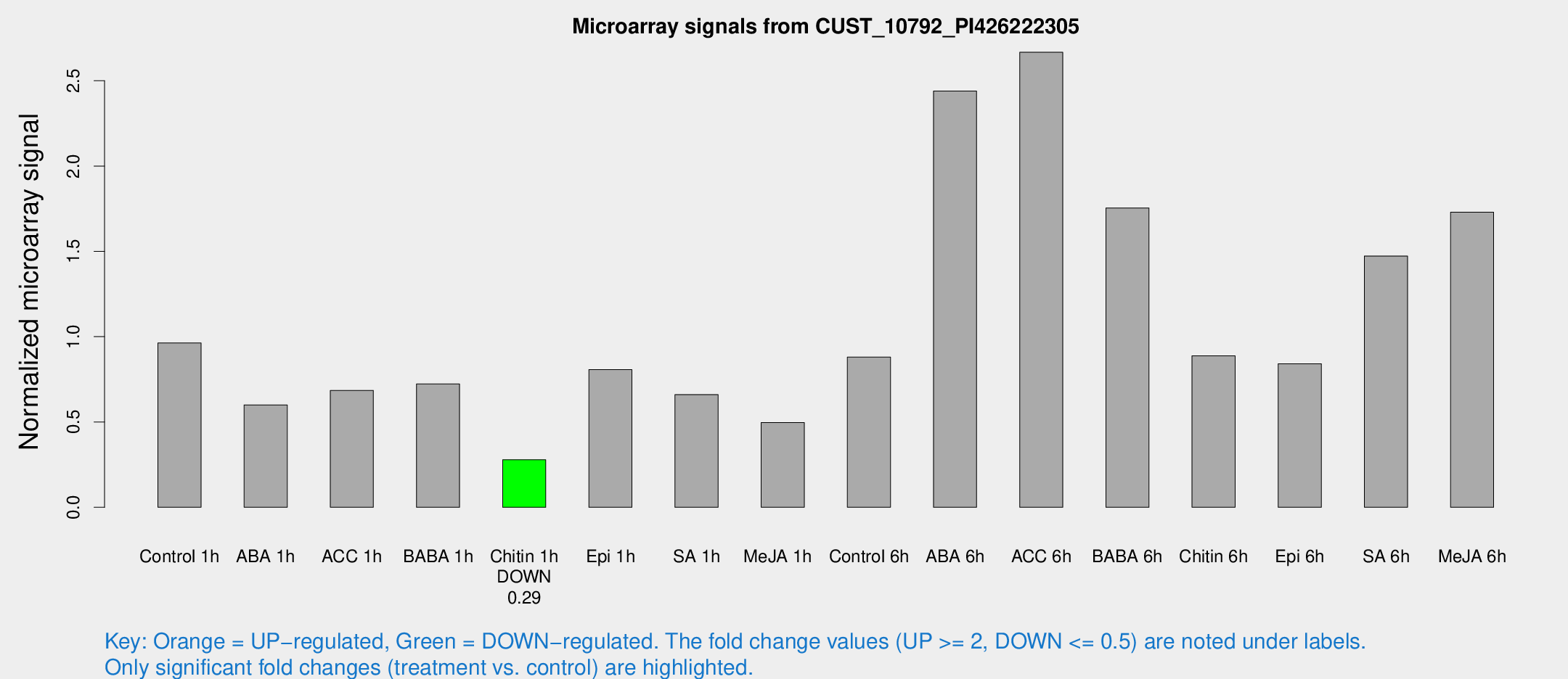
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 34.0431 | 3.98068 | 0.963056 | 0.114295 |
| ABA 1h | 20.2716 | 5.72126 | 0.600248 | 0.228658 |
| ACC 1h | 34.338 | 19.2681 | 0.685169 | 0.448292 |
| BABA 1h | 34.9696 | 15.5106 | 0.722957 | 0.563258 |
| Chitin 1h | 9.14926 | 3.62854 | 0.278651 | 0.121737 |
| Epi 1h | 26.6203 | 7.53634 | 0.807275 | 0.274651 |
| SA 1h | 27.0031 | 8.30641 | 0.660652 | 0.243539 |
| Me-JA 1h | 14.3361 | 3.7954 | 0.496883 | 0.132025 |
| Control 6h | 33.6038 | 9.54422 | 0.880262 | 0.206516 |
| ABA 6h | 94.0523 | 16.2622 | 2.43937 | 0.375773 |
| ACC 6h | 110.215 | 18.9747 | 2.66707 | 0.779653 |
| BABA 6h | 74.5187 | 22.0463 | 1.75451 | 0.444121 |
| Chitin 6h | 33.2053 | 4.4634 | 0.887747 | 0.12041 |
| Epi 6h | 38.497 | 12.5478 | 0.841575 | 0.312399 |
| SA 6h | 52.379 | 8.47351 | 1.47286 | 0.149168 |
| Me-JA 6h | 61.2774 | 7.93711 | 1.72934 | 0.290615 |
Source Transcript PGSC0003DMT400031783 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G63820.1 | +2 | 7e-49 | 171 | 94/166 (57%) | CCT motif family protein | chr1:23682529-23684050 REVERSE LENGTH=293 |
