Probe CUST_10794_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10794_PI426222305 | JHI_St_60k_v1 | DMT400031801 | GAGAATTCATGGTTGATCCTACCTTGAGACGAGTTGAATGAAAACAACTTTATATTGAGG |
All Microarray Probes Designed to Gene DMG400012200
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10794_PI426222305 | JHI_St_60k_v1 | DMT400031801 | GAGAATTCATGGTTGATCCTACCTTGAGACGAGTTGAATGAAAACAACTTTATATTGAGG |
| CUST_10474_PI426222305 | JHI_St_60k_v1 | DMT400031802 | GTGAGAATTCATGGTTGATCCTACCTTGAGACGAGTTGAATGAAAACAACTTTATATTGA |
Microarray Signals from CUST_10794_PI426222305
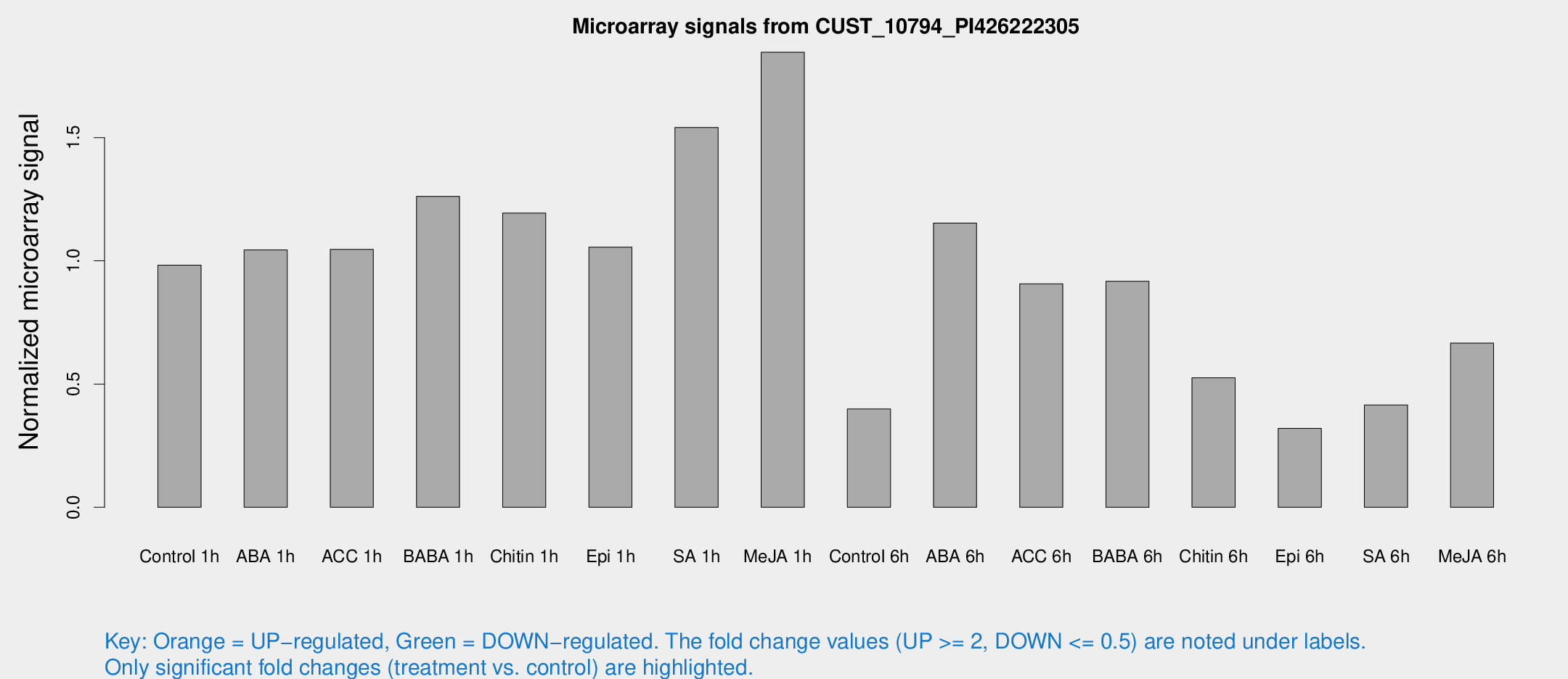
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 49.5004 | 7.4893 | 0.982673 | 0.103326 |
| ABA 1h | 46.8543 | 7.6773 | 1.0444 | 0.152751 |
| ACC 1h | 61.1189 | 19.3682 | 1.04639 | 0.375749 |
| BABA 1h | 61.9651 | 10.4427 | 1.26148 | 0.115804 |
| Chitin 1h | 54.0365 | 8.07639 | 1.19364 | 0.117397 |
| Epi 1h | 45.2327 | 4.42673 | 1.05566 | 0.104106 |
| SA 1h | 82.0294 | 17.7372 | 1.5412 | 0.258078 |
| Me-JA 1h | 74.456 | 5.90785 | 1.84626 | 0.147404 |
| Control 6h | 19.9529 | 4.06903 | 0.398883 | 0.0838618 |
| ABA 6h | 71.117 | 23.1376 | 1.15313 | 0.521344 |
| ACC 6h | 52.8992 | 9.7273 | 0.906577 | 0.278275 |
| BABA 6h | 54.8792 | 15.0889 | 0.917238 | 0.274021 |
| Chitin 6h | 29.7688 | 7.85775 | 0.525847 | 0.147322 |
| Epi 6h | 18.9592 | 4.6906 | 0.32043 | 0.0882276 |
| SA 6h | 25.4022 | 9.83404 | 0.414848 | 0.193637 |
| Me-JA 6h | 33.5948 | 5.64395 | 0.665921 | 0.0874158 |
Source Transcript PGSC0003DMT400031801 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G52430.1 | +1 | 5e-81 | 266 | 206/467 (44%) | hydroxyproline-rich glycoprotein family protein | chr5:21283093-21285045 REVERSE LENGTH=438 |
