Probe CUST_10474_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10474_PI426222305 | JHI_St_60k_v1 | DMT400031802 | GTGAGAATTCATGGTTGATCCTACCTTGAGACGAGTTGAATGAAAACAACTTTATATTGA |
All Microarray Probes Designed to Gene DMG400012200
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10794_PI426222305 | JHI_St_60k_v1 | DMT400031801 | GAGAATTCATGGTTGATCCTACCTTGAGACGAGTTGAATGAAAACAACTTTATATTGAGG |
| CUST_10474_PI426222305 | JHI_St_60k_v1 | DMT400031802 | GTGAGAATTCATGGTTGATCCTACCTTGAGACGAGTTGAATGAAAACAACTTTATATTGA |
Microarray Signals from CUST_10474_PI426222305
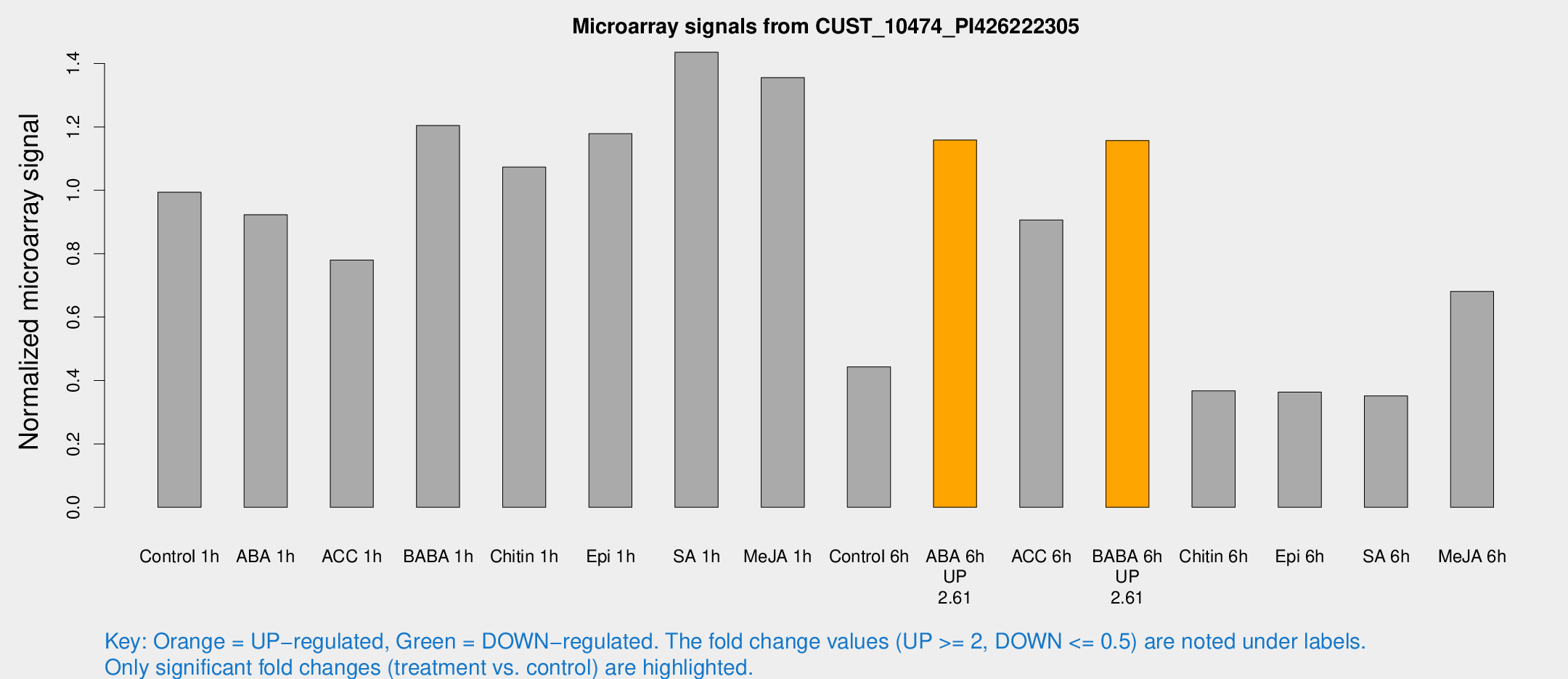
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 53.0667 | 4.57109 | 0.993823 | 0.0863753 |
| ABA 1h | 45.7768 | 9.64287 | 0.922639 | 0.175593 |
| ACC 1h | 49.4709 | 16.0973 | 0.779713 | 0.295682 |
| BABA 1h | 62.7494 | 8.09038 | 1.20433 | 0.1041 |
| Chitin 1h | 51.5819 | 4.72042 | 1.07322 | 0.0981358 |
| Epi 1h | 55.1478 | 6.96996 | 1.17832 | 0.145239 |
| SA 1h | 80.8055 | 13.674 | 1.43542 | 0.172309 |
| Me-JA 1h | 59.5958 | 7.19165 | 1.35531 | 0.117375 |
| Control 6h | 26.5834 | 9.23076 | 0.443059 | 0.12285 |
| ABA 6h | 67.3222 | 11.0685 | 1.15852 | 0.174111 |
| ACC 6h | 56.2259 | 7.59368 | 0.906404 | 0.203483 |
| BABA 6h | 71.1294 | 13.4686 | 1.15708 | 0.171567 |
| Chitin 6h | 20.9795 | 4.32441 | 0.367338 | 0.078027 |
| Epi 6h | 22.1821 | 4.82457 | 0.363258 | 0.0791872 |
| SA 6h | 18.6274 | 4.17766 | 0.351695 | 0.0799781 |
| Me-JA 6h | 37.8682 | 8.42551 | 0.680689 | 0.144527 |
Source Transcript PGSC0003DMT400031802 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G52430.1 | +1 | 2e-79 | 259 | 193/440 (44%) | hydroxyproline-rich glycoprotein family protein | chr5:21283093-21285045 REVERSE LENGTH=438 |
