Probe CUST_10744_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10744_PI426222305 | JHI_St_60k_v1 | DMT400031649 | ATTTAAAAACACCGAACTAAACCGATCCGAAGTAGAAAATATCGAGCCGAACGGATCTAG |
All Microarray Probes Designed to Gene DMG400012137
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10549_PI426222305 | JHI_St_60k_v1 | DMT400031650 | CCTAGTTTTGTTGCATTTCTTGGAATTTGTAAGGCACACAAAAGTGTTGCATATTAGAGT |
| CUST_10744_PI426222305 | JHI_St_60k_v1 | DMT400031649 | ATTTAAAAACACCGAACTAAACCGATCCGAAGTAGAAAATATCGAGCCGAACGGATCTAG |
Microarray Signals from CUST_10744_PI426222305
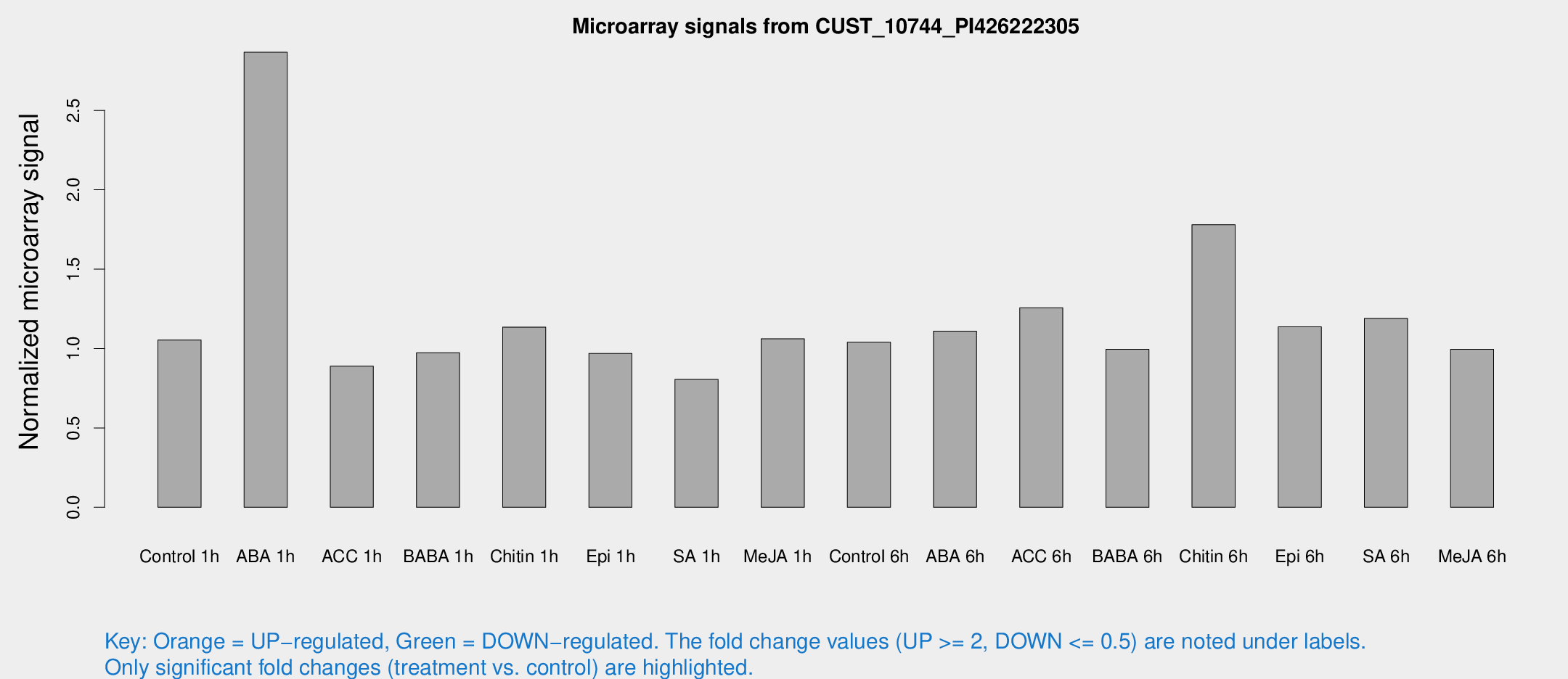
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 7.87501 | 3.53063 | 1.05412 | 0.520241 |
| ABA 1h | 63.9074 | 58.1933 | 2.86639 | 14.7701 |
| ACC 1h | 6.50987 | 3.78118 | 0.889373 | 0.516007 |
| BABA 1h | 6.68934 | 3.75812 | 0.973895 | 0.546389 |
| Chitin 1h | 7.40308 | 3.55102 | 1.13523 | 0.568931 |
| Epi 1h | 5.98661 | 3.43859 | 0.96886 | 0.55573 |
| SA 1h | 5.90284 | 3.4251 | 0.80559 | 0.466643 |
| Me-JA 1h | 6.2007 | 3.60943 | 1.06139 | 0.614732 |
| Control 6h | 7.60605 | 3.78 | 1.03933 | 0.538486 |
| ABA 6h | 8.57393 | 4.00993 | 1.10921 | 0.539715 |
| ACC 6h | 10.3101 | 4.49807 | 1.25653 | 0.552942 |
| BABA 6h | 7.98238 | 4.27124 | 0.995456 | 0.531979 |
| Chitin 6h | 18.0311 | 9.99479 | 1.77975 | 1.1508 |
| Epi 6h | 10.2738 | 4.38974 | 1.13727 | 0.559525 |
| SA 6h | 8.67703 | 4.07297 | 1.18971 | 0.597998 |
| Me-JA 6h | 7.15296 | 3.83046 | 0.994742 | 0.540424 |
Source Transcript PGSC0003DMT400031649 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G24015.1 | +1 | 2e-24 | 97 | 45/79 (57%) | RING/U-box superfamily protein | chr4:12469887-12471197 REVERSE LENGTH=174 |
