Probe CUST_10549_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10549_PI426222305 | JHI_St_60k_v1 | DMT400031650 | CCTAGTTTTGTTGCATTTCTTGGAATTTGTAAGGCACACAAAAGTGTTGCATATTAGAGT |
All Microarray Probes Designed to Gene DMG400012137
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10549_PI426222305 | JHI_St_60k_v1 | DMT400031650 | CCTAGTTTTGTTGCATTTCTTGGAATTTGTAAGGCACACAAAAGTGTTGCATATTAGAGT |
| CUST_10744_PI426222305 | JHI_St_60k_v1 | DMT400031649 | ATTTAAAAACACCGAACTAAACCGATCCGAAGTAGAAAATATCGAGCCGAACGGATCTAG |
Microarray Signals from CUST_10549_PI426222305
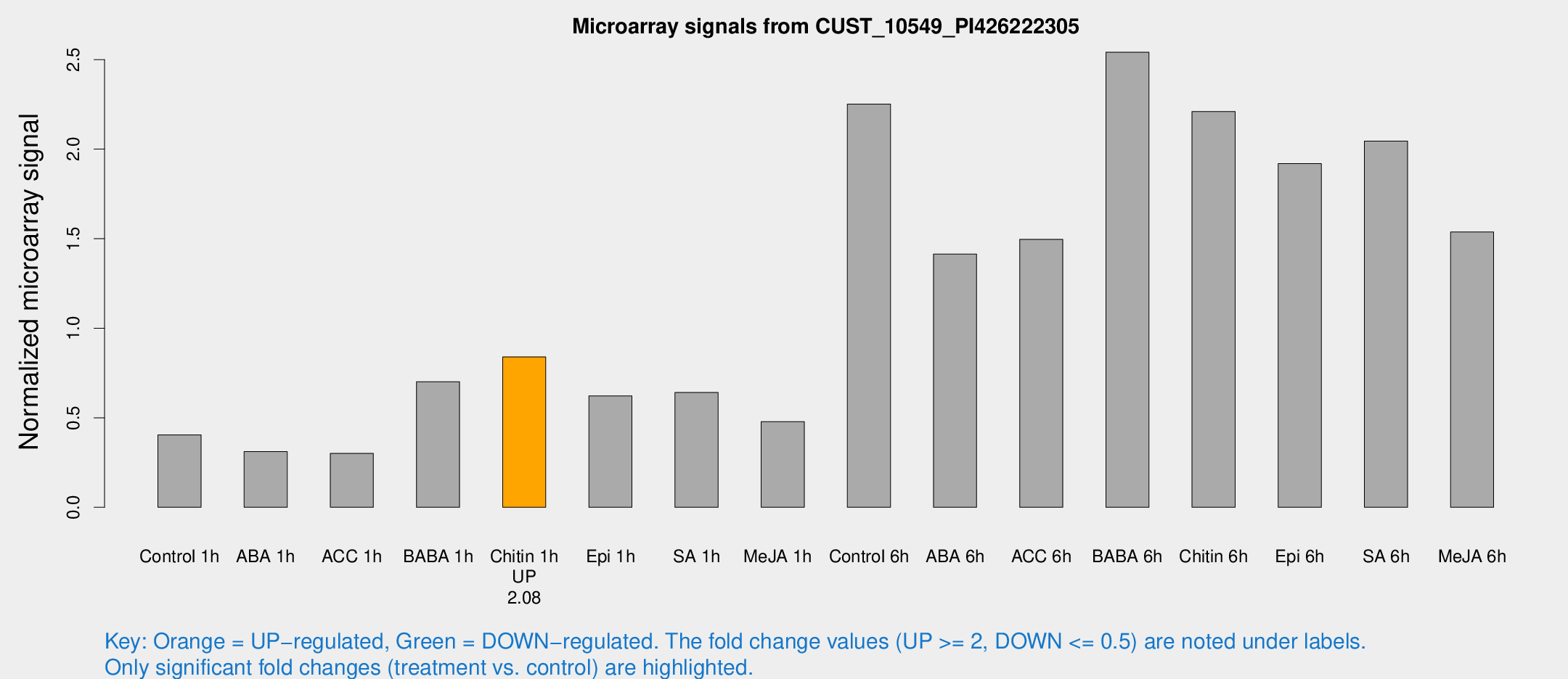
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 45.3259 | 6.93511 | 0.404112 | 0.0392934 |
| ABA 1h | 30.9772 | 4.83658 | 0.310987 | 0.0398531 |
| ACC 1h | 42.5251 | 16.3753 | 0.300883 | 0.157292 |
| BABA 1h | 76.8157 | 13.8974 | 0.701401 | 0.0700574 |
| Chitin 1h | 82.9075 | 5.90969 | 0.839543 | 0.0597282 |
| Epi 1h | 59.6151 | 5.68631 | 0.622026 | 0.0509094 |
| SA 1h | 73.9333 | 10.8495 | 0.640861 | 0.122208 |
| Me-JA 1h | 46.7404 | 12.3102 | 0.478684 | 0.11329 |
| Control 6h | 272.06 | 72.4295 | 2.25131 | 0.552425 |
| ABA 6h | 168.318 | 25.0369 | 1.41353 | 0.134107 |
| ACC 6h | 200.848 | 48.2496 | 1.49621 | 0.23991 |
| BABA 6h | 312.281 | 18.4905 | 2.54109 | 0.150414 |
| Chitin 6h | 261.481 | 30.1771 | 2.20972 | 0.181671 |
| Epi 6h | 237.923 | 14.3763 | 1.91927 | 0.209652 |
| SA 6h | 225.069 | 27.3977 | 2.04454 | 0.193533 |
| Me-JA 6h | 178.521 | 40.5761 | 1.53734 | 0.310047 |
Source Transcript PGSC0003DMT400031650 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G24015.1 | +1 | 3e-37 | 134 | 96/154 (62%) | RING/U-box superfamily protein | chr4:12469887-12471197 REVERSE LENGTH=174 |
