Probe CUST_10729_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10729_PI426222305 | JHI_St_60k_v1 | DMT400031808 | CTCCCGATATTTTTGACTATTAGCTTATTTTGGTTCTCCAGCAATCACCATGCTAGGAGC |
All Microarray Probes Designed to Gene DMG400012203
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10729_PI426222305 | JHI_St_60k_v1 | DMT400031808 | CTCCCGATATTTTTGACTATTAGCTTATTTTGGTTCTCCAGCAATCACCATGCTAGGAGC |
| CUST_10525_PI426222305 | JHI_St_60k_v1 | DMT400031805 | AAGGAGAAGGCTCTCTCAGTATCCCAATCTATTCGTTCCCAGTTGCTATCCAATCACCAT |
Microarray Signals from CUST_10729_PI426222305
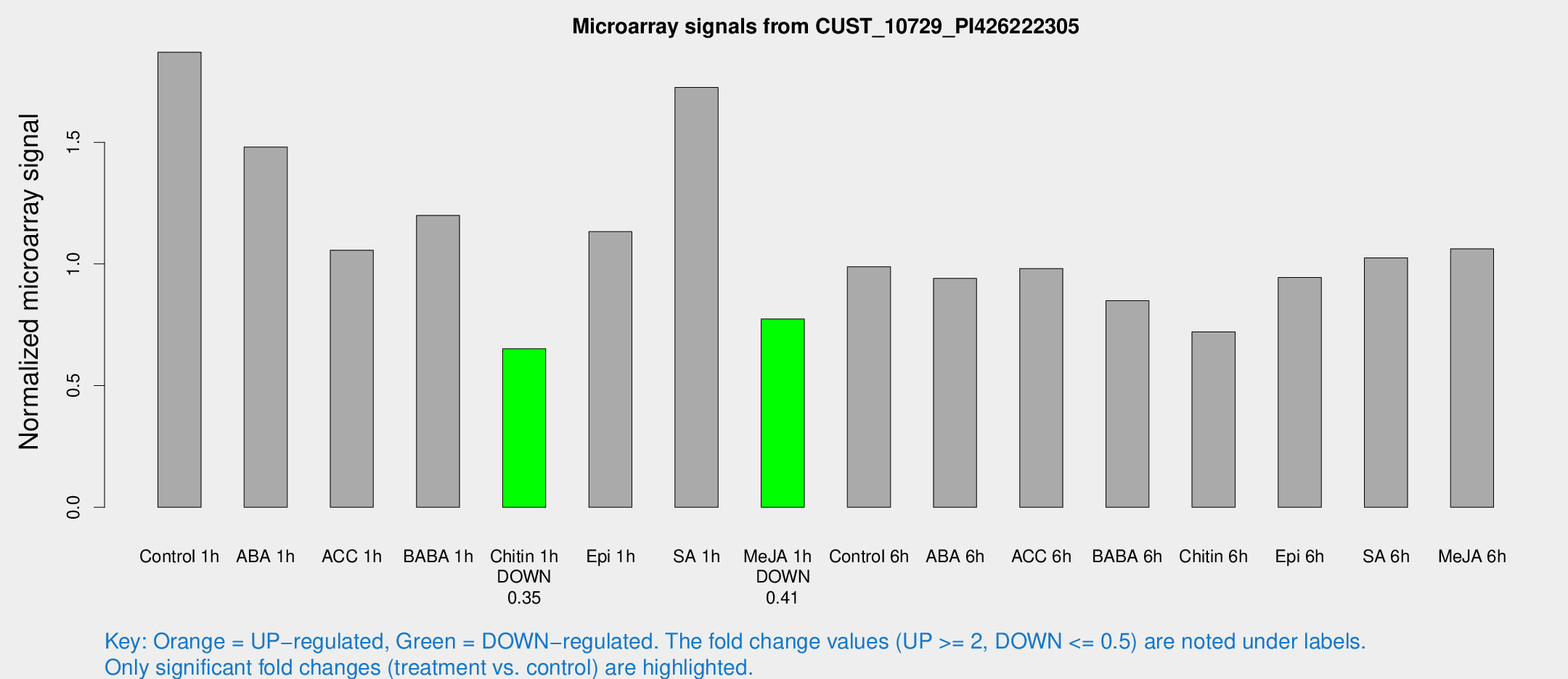
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 75.5105 | 13.1756 | 1.87005 | 0.230288 |
| ABA 1h | 52.7584 | 8.9183 | 1.48049 | 0.167405 |
| ACC 1h | 46.5869 | 12.5283 | 1.05661 | 0.278034 |
| BABA 1h | 49.1224 | 13.2427 | 1.19936 | 0.250793 |
| Chitin 1h | 24.6879 | 6.01882 | 0.651486 | 0.157959 |
| Epi 1h | 39.3762 | 6.16749 | 1.13269 | 0.177363 |
| SA 1h | 69.9166 | 7.11633 | 1.72541 | 0.189349 |
| Me-JA 1h | 24.8308 | 3.91035 | 0.773866 | 0.120703 |
| Control 6h | 45.8663 | 15.528 | 0.988704 | 0.393043 |
| ABA 6h | 39.5982 | 4.51373 | 0.940813 | 0.107804 |
| ACC 6h | 44.2708 | 5.06885 | 0.981346 | 0.212162 |
| BABA 6h | 37.5794 | 4.71406 | 0.849016 | 0.10805 |
| Chitin 6h | 30.3561 | 4.61013 | 0.720766 | 0.112303 |
| Epi 6h | 42.8659 | 6.96139 | 0.944505 | 0.116953 |
| SA 6h | 48.5219 | 22.4032 | 1.02505 | 0.413058 |
| Me-JA 6h | 45.0473 | 11.6259 | 1.06199 | 0.256358 |
Source Transcript PGSC0003DMT400031808 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G63700.1 | +3 | 2e-13 | 73 | 72/190 (38%) | Protein kinase superfamily protein | chr1:23625208-23629031 REVERSE LENGTH=883 |
