Probe CUST_10525_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10525_PI426222305 | JHI_St_60k_v1 | DMT400031805 | AAGGAGAAGGCTCTCTCAGTATCCCAATCTATTCGTTCCCAGTTGCTATCCAATCACCAT |
All Microarray Probes Designed to Gene DMG400012203
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10729_PI426222305 | JHI_St_60k_v1 | DMT400031808 | CTCCCGATATTTTTGACTATTAGCTTATTTTGGTTCTCCAGCAATCACCATGCTAGGAGC |
| CUST_10525_PI426222305 | JHI_St_60k_v1 | DMT400031805 | AAGGAGAAGGCTCTCTCAGTATCCCAATCTATTCGTTCCCAGTTGCTATCCAATCACCAT |
Microarray Signals from CUST_10525_PI426222305
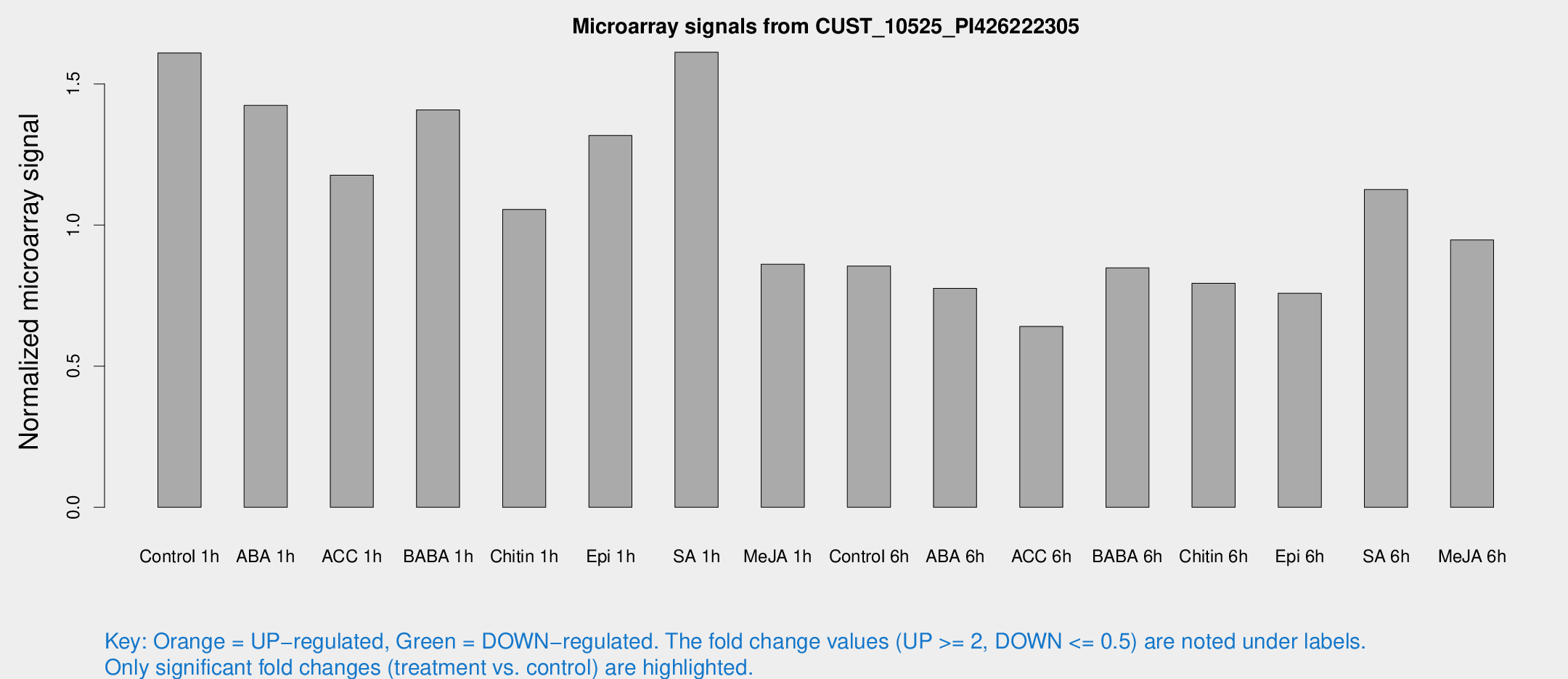
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 417.447 | 60.5958 | 1.60961 | 0.126314 |
| ABA 1h | 321.48 | 19.7059 | 1.42387 | 0.0836341 |
| ACC 1h | 355.376 | 111.697 | 1.17653 | 0.432216 |
| BABA 1h | 360.073 | 70.7858 | 1.40817 | 0.163932 |
| Chitin 1h | 246.57 | 34.2874 | 1.05536 | 0.106844 |
| Epi 1h | 296.174 | 40.5683 | 1.31765 | 0.178642 |
| SA 1h | 425.828 | 45.4675 | 1.6124 | 0.166027 |
| Me-JA 1h | 180.459 | 18.207 | 0.860947 | 0.0527945 |
| Control 6h | 236.161 | 67.2425 | 0.854761 | 0.198765 |
| ABA 6h | 211.075 | 16.8914 | 0.775961 | 0.10436 |
| ACC 6h | 186.902 | 11.7014 | 0.640663 | 0.0903678 |
| BABA 6h | 243.678 | 23.1957 | 0.84828 | 0.0625562 |
| Chitin 6h | 221.283 | 39.3971 | 0.793988 | 0.137353 |
| Epi 6h | 228.448 | 46.1782 | 0.758647 | 0.122801 |
| SA 6h | 302.306 | 80.6551 | 1.12599 | 0.222316 |
| Me-JA 6h | 246.75 | 38.7489 | 0.947482 | 0.103264 |
Source Transcript PGSC0003DMT400031805 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G63700.1 | +1 | 0.0 | 764 | 494/888 (56%) | Protein kinase superfamily protein | chr1:23625208-23629031 REVERSE LENGTH=883 |
