Probe CUST_10358_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10358_PI426222305 | JHI_St_60k_v1 | DMT400029078 | GCGTTTGGCTGTTTGTCTAGATCCTGATCACATGCCACTATTTGAAAAGTATTTGTATGA |
All Microarray Probes Designed to Gene DMG400011189
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10358_PI426222305 | JHI_St_60k_v1 | DMT400029078 | GCGTTTGGCTGTTTGTCTAGATCCTGATCACATGCCACTATTTGAAAAGTATTTGTATGA |
| CUST_10321_PI426222305 | JHI_St_60k_v1 | DMT400029079 | CTCATTCCTAGTGTTGAAACTTCTGGAGATATCTCAACCTTTCCACTCGCTATATTTCAG |
Microarray Signals from CUST_10358_PI426222305
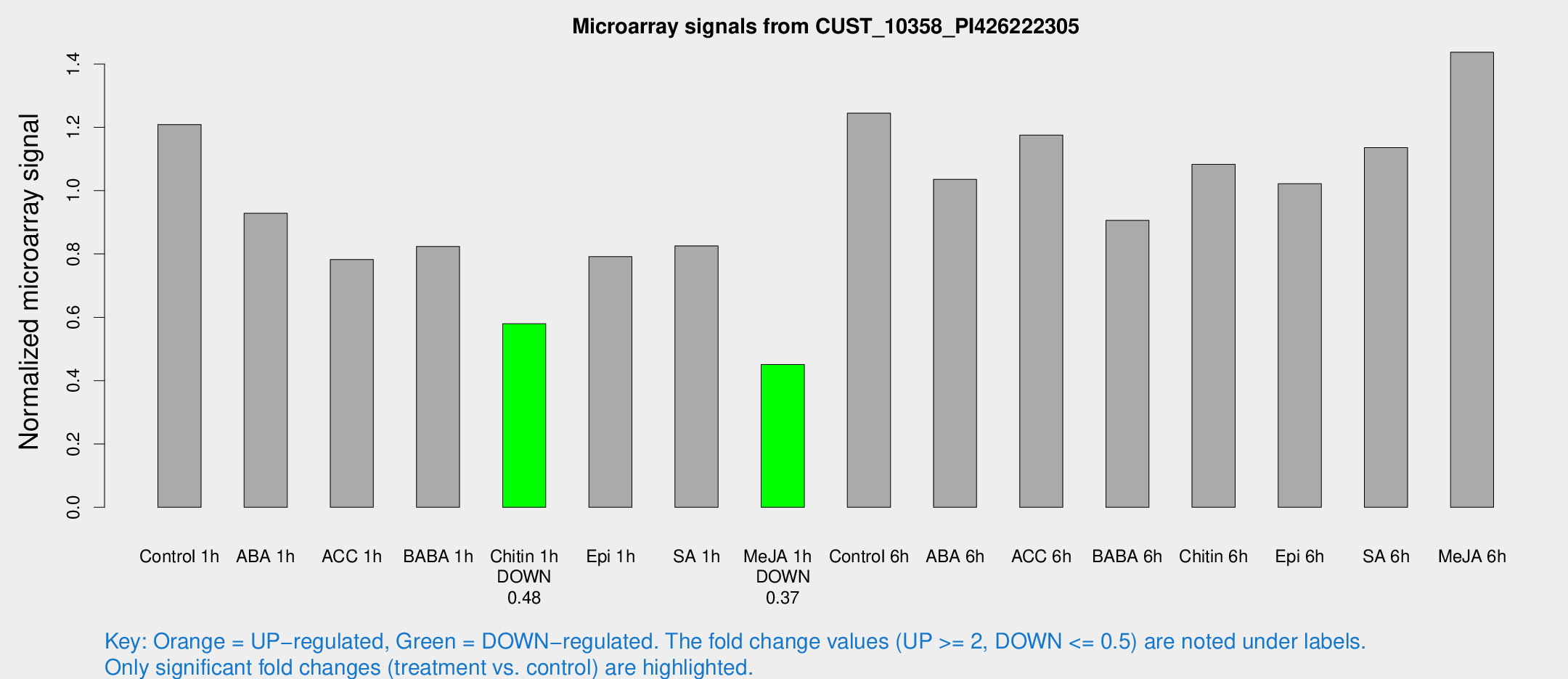
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 26383.2 | 2529.2 | 1.20881 | 0.0697907 |
| ABA 1h | 17839.6 | 1270.6 | 0.92824 | 0.068047 |
| ACC 1h | 21205.3 | 7405.23 | 0.782268 | 0.376784 |
| BABA 1h | 17930.4 | 3473.65 | 0.823519 | 0.0944812 |
| Chitin 1h | 11653.1 | 2232.81 | 0.579684 | 0.0680628 |
| Epi 1h | 14902 | 1199.31 | 0.791601 | 0.0704271 |
| SA 1h | 19827.5 | 5364.19 | 0.825495 | 0.220731 |
| Me-JA 1h | 8059.44 | 922.59 | 0.450789 | 0.026027 |
| Control 6h | 27478 | 3622.2 | 1.24453 | 0.0772822 |
| ABA 6h | 23985.2 | 2140.71 | 1.03576 | 0.0598001 |
| ACC 6h | 29153.4 | 1683.27 | 1.1756 | 0.164675 |
| BABA 6h | 22081.3 | 1915.77 | 0.906015 | 0.0720252 |
| Chitin 6h | 25128.7 | 2197.33 | 1.0832 | 0.131145 |
| Epi 6h | 24923.1 | 1440.27 | 1.02166 | 0.0863622 |
| SA 6h | 25344.9 | 4892.97 | 1.13578 | 0.113292 |
| Me-JA 6h | 31647.7 | 4938.06 | 1.43703 | 0.161248 |
Source Transcript PGSC0003DMT400029078 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G48930.1 | +2 | 1e-117 | 351 | 167/297 (56%) | hydroxycinnamoyl-CoA shikimate/quinate hydroxycinnamoyl transferase | chr5:19836654-19838092 REVERSE LENGTH=433 |
