Probe CUST_10321_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10321_PI426222305 | JHI_St_60k_v1 | DMT400029079 | CTCATTCCTAGTGTTGAAACTTCTGGAGATATCTCAACCTTTCCACTCGCTATATTTCAG |
All Microarray Probes Designed to Gene DMG400011189
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10358_PI426222305 | JHI_St_60k_v1 | DMT400029078 | GCGTTTGGCTGTTTGTCTAGATCCTGATCACATGCCACTATTTGAAAAGTATTTGTATGA |
| CUST_10321_PI426222305 | JHI_St_60k_v1 | DMT400029079 | CTCATTCCTAGTGTTGAAACTTCTGGAGATATCTCAACCTTTCCACTCGCTATATTTCAG |
Microarray Signals from CUST_10321_PI426222305
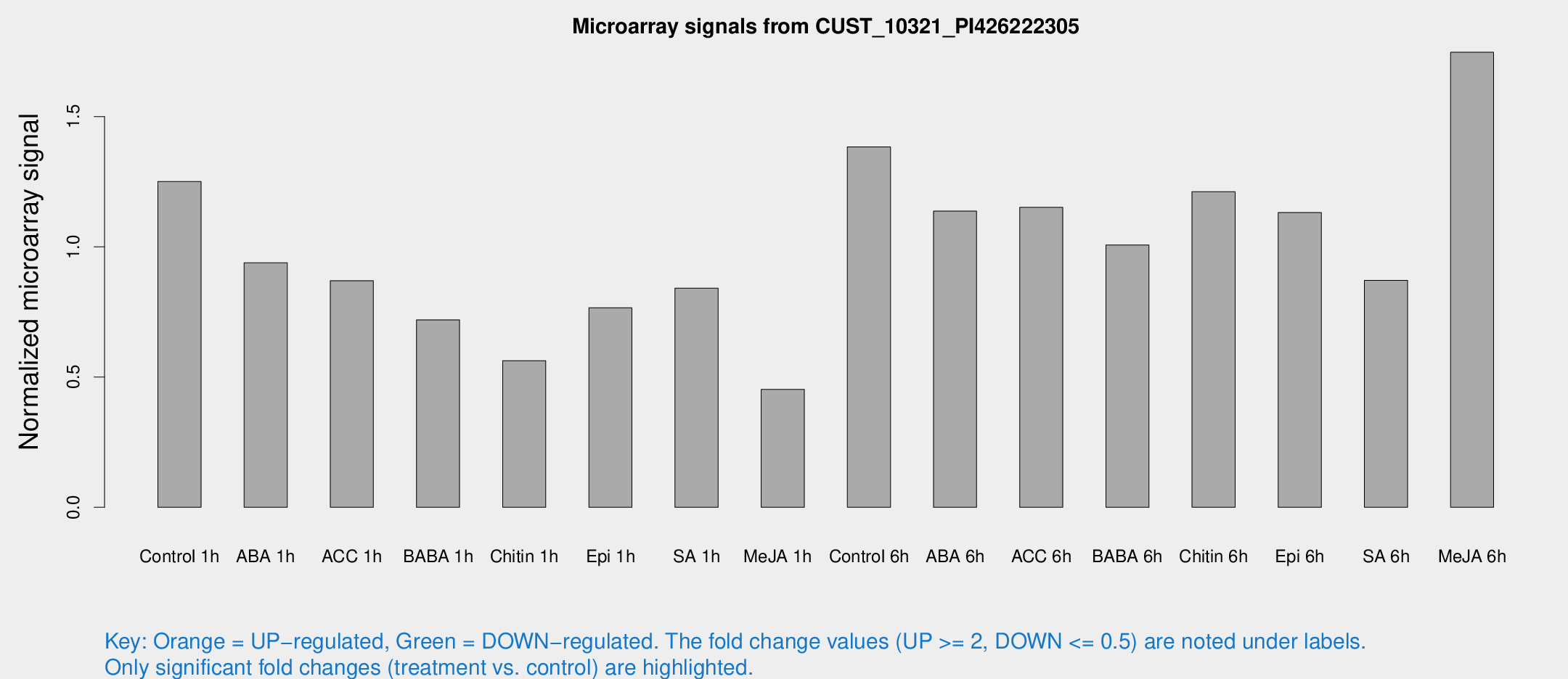
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 3322.47 | 567.133 | 1.25094 | 0.132004 |
| ABA 1h | 2166.44 | 227.776 | 0.938711 | 0.0542105 |
| ACC 1h | 2836.07 | 1009.7 | 0.869941 | 0.43153 |
| BABA 1h | 1942.64 | 488.274 | 0.719537 | 0.147351 |
| Chitin 1h | 1384.95 | 353.21 | 0.562898 | 0.0964942 |
| Epi 1h | 1717.52 | 108.869 | 0.766187 | 0.0676807 |
| SA 1h | 2474.44 | 691.236 | 0.841527 | 0.267317 |
| Me-JA 1h | 981.367 | 163.184 | 0.452503 | 0.0406125 |
| Control 6h | 3981.16 | 1206 | 1.38347 | 0.379849 |
| ABA 6h | 3219.06 | 541.629 | 1.13697 | 0.153684 |
| ACC 6h | 3428.82 | 269.711 | 1.15184 | 0.0719845 |
| BABA 6h | 3016.61 | 596.02 | 1.00719 | 0.179381 |
| Chitin 6h | 3340.94 | 226.442 | 1.21142 | 0.1132 |
| Epi 6h | 3389.39 | 605.471 | 1.13169 | 0.202462 |
| SA 6h | 2655.09 | 925.288 | 0.871144 | 0.326585 |
| Me-JA 6h | 4605.96 | 759.71 | 1.74718 | 0.145829 |
Source Transcript PGSC0003DMT400029079 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G48930.1 | +1 | 0.0 | 524 | 248/433 (57%) | hydroxycinnamoyl-CoA shikimate/quinate hydroxycinnamoyl transferase | chr5:19836654-19838092 REVERSE LENGTH=433 |
