- Transcript BART1_0-u54453.008
- Transcript IDBART1_0-u54453.008
- Gene IDBART1_0-u54453
Exon Structure of BART1_0-u54453.008
| Chromosome | Exon | Start | Stop | Direction |
|---|---|---|---|---|
| chr7H | 1 | 514441144 | 514440918 | - |
| chr7H | 2 | 514442287 | 514442135 | - |
| chr7H | 3 | 514442474 | 514442357 | - |
| chr7H | 4 | 514442603 | 514442556 | - |
| chr7H | 5 | 514442934 | 514442719 | - |
| chr7H | 6 | 514443079 | 514443015 | - |
| chr7H | 7 | 514443422 | 514443298 | - |
| chr7H | 8 | 514443642 | 514443512 | - |
| chr7H | 9 | 514444758 | 514444470 | - |
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
Salmon TPM Values: sixteensamplesmorex
These TPM values were generated by using the RNA-seq data from a 16-tissue experiment in Morex (published here) were calculated using Salmon (version Salmon-0.8.2) using BaRTv1.0-QUASI as the reference transcript dataset. Click here for more information about the RNA-seq experiment and materials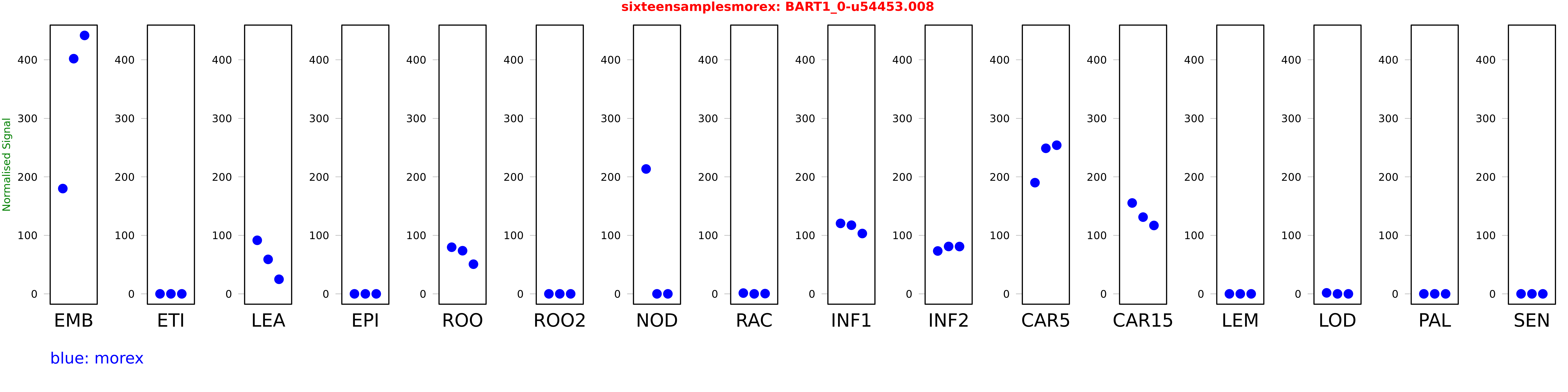
| Sample | Treatment | Rep 1 | Rep 2 | Rep 3 |
|---|---|---|---|---|
| morex | EMB | 179.953 | 401.961 | 441.707 |
| morex | ETI | 0 | 0 | 0 |
| morex | LEA | 91.5476 | 58.9369 | 24.9228 |
| morex | EPI | 0 | 0 | 0 |
| morex | ROO | 79.7018 | 73.7202 | 50.6903 |
| morex | ROO2 | 0 | 0 | 0 |
| morex | NOD | 213.449 | 0 | 0 |
| morex | RAC | 1.3225 | 0 | 0.425456 |
| morex | INF1 | 120.44 | 117.343 | 103.135 |
| morex | INF2 | 73.2849 | 81.0128 | 80.8759 |
| morex | CAR5 | 190.044 | 248.787 | 254.048 |
| morex | CAR15 | 155.327 | 131.154 | 116.903 |
| morex | LEM | 0 | 0 | 0 |
| morex | LOD | 1.70679 | 0 | 0 |
| morex | PAL | 0 | 0 | 0 |
| morex | SEN | 0 | 0 | 0 |
Homology to Model Species (BLASTX to E-value < 1e-30)
| Database | Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Rice PP7 | LOC_Os08g09250.2 | +2 | 3e-154 | 440 | 210/231 (91%) | protein|glyoxalase family protein, putative, expressed |
| TAIR PP10 | AT1G67280 @ ARAPORT AT1G67280.1 @ TAIR |
+2 | 4e-123 | 362 | 180/261 (69%) | Symbols: | Glyoxalase/Bleomycin resistance protein/Dioxygenase superfamily protein | chr1:25188563-25190547 REVERSE LENGTH=350 |
| BRACH PP3 | Bradi3g14960.1.p | +2 | 1e-158 | 447 | 213/231 (92%) | pacid=32813745 transcript=Bradi3g14960.1 locus=Bradi3g14960 ID=Bradi3g14960.1.v3.1 annot-version=v3.1 |
Barley PseudoMolecules GBrowse
Click here to see more tracks within GBrowse -->
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
CDS Sequence (1372 bp)
>BART1_0-u54453.008 1372 150831_barley_pseudomolecules ATGGCCCATTTGTAACCTTGCTGGGCCAGGCCCATCCATGCCGACACGCGCCCACAGCCC ATCTGGCCAGCTTTAGTGGGAGGGATGCGCTTTTACAGCCTGTGAGGAGGTACCACGTCG CCATCAACGCAATCATAAGTATCGGTGCCGGACTCTCTTCTCCTTGGAAGCCTCTAGACT GAGCTCCTCATCAGCGTGGGACGGATCTCTGCCGGATCGTATTTCCGGGGTGAGCATCCC CAAATCCATCTCCTTCGTCCTCTCTTCCGCTTGCTTCCTTTACGCGCCGGTGACAACAGG GATGGCTACCGGTAGCGAAGCTGGAAAGTCCGCCGAGGCCGTGCTGGAATGGCCTAAGCA GGACAAAAAGAGGATGCTGCATGCTGTTTACCGTGTGGGAGATCTTGACAAAACCATTAA GTGTTACACAGAATGCTTTGGGATGAAGCTGCTGAGGAAAAGAGATGTCCCAGAAGAGAA GTACACCAATGCGTTTCTTGGGTTTGGACCTGAGGATACTAATTTTGCACTTGAGCTGAC TTACAATTATGGTGTTGACAAGTACGACATTGGAGCGGGCTTTGGACATTTTGCCATCGC AAATGAGGATGTGTACAAGCTGTCTGAGACAATTAAATCATCTGATTGTTGTAAGATCAC TCGTGAACCTGGTCCTGTCAAGGGAGGGTCCACTGTGATTGCCTTTGCACAAGACCCAGA TGGTTACTTGTTTGAGCTTATCCAGAGGGGTCCGACGCCTGAGCCTCTCTGTCAAGTTAT GCTTCGTGTTGGTGACCTTGATCGGGCTATCATGTTCTACGAGAAGGCCCTTGGGATGAA GCTTCTGAGGAAGAAGGATGTGCCTCAGTATAAGTACACCATTGCCATGATGGGCTATGC TGAGGAGGACAAGACCACTGTTCTGGAGTTGACATACAACTATGGTGTCACGGAATATAA CAAGGGCAATGCATATGCTCAGGTATTGACAGGTTGCTATTGGCACTGACGATGTGTACA AGAGCGCCGAAGCAGTTGAGCTGGTTACCAAAGAACTAGGTGGAAAGATTCTAAGGCAGC CAGGGCCACTACCGGGGCTGAACACCAAAATCACCTCTTTCCTTGACCCAGATGGCTGGA AAGTGGTTCTGGTGGACTACGCAGACTTCCTCAAGGAGCTGCACTGAAGATGAGCAGGCT CAGATGTTAGTGTCGTGAGGAGGAACCGTATGTAGCAATGGTCTGTGTGATGTAATAAAA CAAACCTCATGTAGCAAATGGTCTGTGTGCGATGTAATAAAAAGGCAGCCTCTTCCGCTC GACGAATAAAAATAAAAGCAACTACCGCATGCCCATGCCGTCATTNNNNNNN
Protein Sequence (235 aa)
>BART1_0-u54453.008 235 150831_barley_pseudomolecules MATGSEAGKSAEAVLEWPKQDKKRMLHAVYRVGDLDKTIKCYTECFGMKLLRKRDVPEEK YTNAFLGFGPEDTNFALELTYNYGVDKYDIGAGFGHFAIANEDVYKLSETIKSSDCCKIT REPGPVKGGSTVIAFAQDPDGYLFELIQRGPTPEPLCQVMLRVGDLDRAIMFYEKALGMK LLRKKDVPQYKYTIAMMGYAEEDKTTVLELTYNYGVTEYNKGNAYAQVLTGCYWH
