- Transcript BART1_0-u54453.007
- Transcript IDBART1_0-u54453.007
- Gene IDBART1_0-u54453
Exon Structure of BART1_0-u54453.007
| Chromosome | Exon | Start | Stop | Direction |
|---|---|---|---|---|
| chr7H | 1 | 514441144 | 514440918 | - |
| chr7H | 2 | 514442287 | 514442135 | - |
| chr7H | 3 | 514442474 | 514442367 | - |
| chr7H | 4 | 514442934 | 514442556 | - |
| chr7H | 5 | 514443079 | 514443015 | - |
| chr7H | 6 | 514443422 | 514443298 | - |
| chr7H | 7 | 514443642 | 514443512 | - |
| chr7H | 8 | 514444206 | 514444065 | - |
| chr7H | 9 | 514444648 | 514444530 | - |
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
Salmon TPM Values: sixteensamplesmorex
These TPM values were generated by using the RNA-seq data from a 16-tissue experiment in Morex (published here) were calculated using Salmon (version Salmon-0.8.2) using BaRTv1.0-QUASI as the reference transcript dataset. Click here for more information about the RNA-seq experiment and materials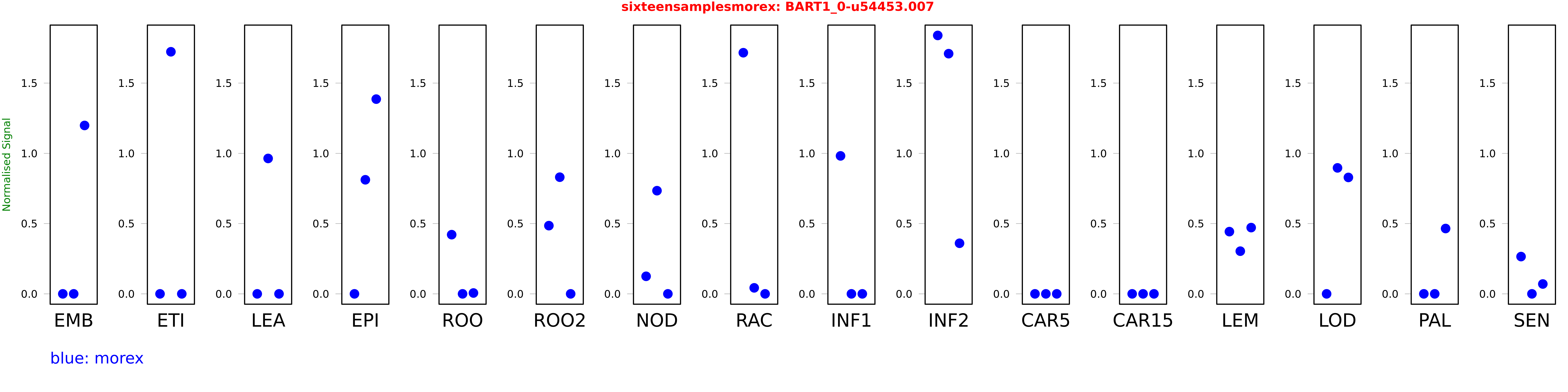
| Sample | Treatment | Rep 1 | Rep 2 | Rep 3 |
|---|---|---|---|---|
| morex | EMB | 0 | 0 | 1.1986 |
| morex | ETI | 0 | 1.72306 | 0 |
| morex | LEA | 0 | 0.963794 | 0 |
| morex | EPI | 0 | 0.81195 | 1.38606 |
| morex | ROO | 0.421242 | 0 | 0.0061297 |
| morex | ROO2 | 0.485497 | 0.830374 | 0 |
| morex | NOD | 0.124956 | 0.73427 | 0 |
| morex | RAC | 1.71629 | 0.0427391 | 0 |
| morex | INF1 | 0.981939 | 0 | 0.000000155712 |
| morex | INF2 | 1.83938 | 1.70948 | 0.360113 |
| morex | CAR5 | 0 | 0 | 0 |
| morex | CAR15 | 0 | 0 | 0 |
| morex | LEM | 0.442905 | 0.303806 | 0.471577 |
| morex | LOD | 0 | 0.896647 | 0.828683 |
| morex | PAL | 0 | 0 | 0.465002 |
| morex | SEN | 0.265325 | 0 | 0.0701971 |
Homology to Model Species (BLASTX to E-value < 1e-30)
| Database | Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Rice PP7 | LOC_Os08g09250.2 | +1 | 5e-153 | 342 | 163/181 (90%) | protein|glyoxalase family protein, putative, expressed |
| TAIR PP10 | AT1G11840 @ ARAPORT AT1G11840.6 @ TAIR |
+1 | 7e-120 | 268 | 123/171 (72%) | Symbols: GLX1 | glyoxalase I homolog | chr1:3995928-3997518 FORWARD LENGTH=322 |
| BRACH PP3 | Bradi3g14960.1.p | +1 | 1e-157 | 348 | 165/181 (91%) | pacid=32813745 transcript=Bradi3g14960.1 locus=Bradi3g14960 ID=Bradi3g14960.1.v3.1 annot-version=v3.1 |
Barley PseudoMolecules GBrowse
Click here to see more tracks within GBrowse -->
grey: non-coding, green: Barley Morex IBSC 2017 Transcript CDS, red: Barley RTD exons
CDS Sequence (1449 bp)
>BART1_0-u54453.007 1449 150831_barley_pseudomolecules TACCACGTCGCCATCAACGCAATCATAAGTATCGGTGCCGGACTCTCTTCTCCTTGGAAG CCTCTAGACTGAGCTCCTCATCAGCGTGGGACGGATCTCTGCCGGATCGTATTTCCGGGC AGAACATCTTCCTGGCGTGGTAGCGGCTGGGCTGTGGGGGAGGTGCTCATGGATGGATTG GGATCCTTTCGCCTCAGCGCAGGGATTTGGGGGCGCTACTTATTTATCCAGTCCTGCCGC TAGTTGCTGCTAATTTATCCAGTGACAACAGGGATGGCTACCGGTAGCGAAGCTGGAAAG TCCGCCGAGGCCGTGCTGGAATGGCCTAAGCAGGACAAAAAGAGGATGCTGCATGCTGTT TACCGTGTGGGAGATCTTGACAAAACCATTAAGTGTTACACAGAATGCTTTGGGATGAAG CTGCTGAGGAAAAGAGATGTCCCAGAAGAGAAGTACACCAATGCGTTTCTTGGGTTTGGA CCTGAGGATACTAATTTTGCACTTGAGCTGACTTACAATTATGGTGTTGACAAGTACGAC ATTGGAGCGGGCTTTGGACATTTTGCCATCGCAAATGAGGATGTGTACAAGCTGTCTGAG ACAATTAAATCATCTGATTGTTGTAAGATCACTCGTGAACCTGGTCCTGTCAAGGGAGGG TCCACTGTGATTGCCTTTGCACAAGACCCAGATGGTTACTTGTTTGAGCTTATCCAGAGG GGTCCGACGCCTGAGCCTCTCTGTCAAGTTATGCTTCGTGTTGGTGACCTTGATCGGGCT ATCATGTTCTACGAGAAGGTATTGTTTTGTAAAATGGGAGGTTGTTTGAAATGTTAAGAG ACAGTATCCTTTCTGTATATTCTCTAGAACTATAGTTTTATATGGGATATTCTAACATTT GACTTTTATGCAGGCCCTTGGGATGAAGCTTCTGAGGAAGAAGGATGTGCCTCAGTATAA GTACACCATTGCCATGATGGGCTATGCTGAGGAGGACAAGACCACTGTTCTGGAGTTGAC ATACAACTATGGTGTCACGGAATATAACAAGGGCAATGCATATGCTCAGGTTGCTATTGG CACTGACGATGTGTACAAGAGCGCCGAAGCAGTTGAGCTGGTTACCAAAGAACTAGGTGG AAAGATTCTAAGGCAGCCAGGGCCACTACCGGGGCTGAACACCAAAATCACCTCTTTCCT TGACCCAGATGGCTGGAAAGTGGTTCTGGTGGACTACGCAGACTTCCTCAAGGAGCTGCA CTGAAGATGAGCAGGCTCAGATGTTAGTGTCGTGAGGAGGAACCGTATGTAGCAATGGTC TGTGTGATGTAATAAAACAAACCTCATGTAGCAAATGGTCTGTGTGCGATGTAATAAAAA GGCAGCCTCTTCCGCTCGACGAATAAAAATAAAAGCAACTACCGCATGCCCATGCCGTCA TTNNNNNNN
Protein Sequence (187 aa)
>BART1_0-u54453.007 187 150831_barley_pseudomolecules MATGSEAGKSAEAVLEWPKQDKKRMLHAVYRVGDLDKTIKCYTECFGMKLLRKRDVPEEK YTNAFLGFGPEDTNFALELTYNYGVDKYDIGAGFGHFAIANEDVYKLSETIKSSDCCKIT REPGPVKGGSTVIAFAQDPDGYLFELIQRGPTPEPLCQVMLRVGDLDRAIMFYEKVLFCK MGGCLKC
