Probe CUST_8723_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_8723_PI426222305 | JHI_St_60k_v1 | DMT400071690 | AACCATGGGTGCAATGGAGGGCTTCCATCACATGCGTTTGAGTACATTAAATACAATGGT |
All Microarray Probes Designed to Gene DMG400027888
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_8723_PI426222305 | JHI_St_60k_v1 | DMT400071690 | AACCATGGGTGCAATGGAGGGCTTCCATCACATGCGTTTGAGTACATTAAATACAATGGT |
| CUST_8792_PI426222305 | JHI_St_60k_v1 | DMT400071689 | TGGCCTTAGGCCCCCAATTTTAGAGGAGCCCATTTTTTAGTTAATTGGGTAAGATTAAAT |
Microarray Signals from CUST_8723_PI426222305
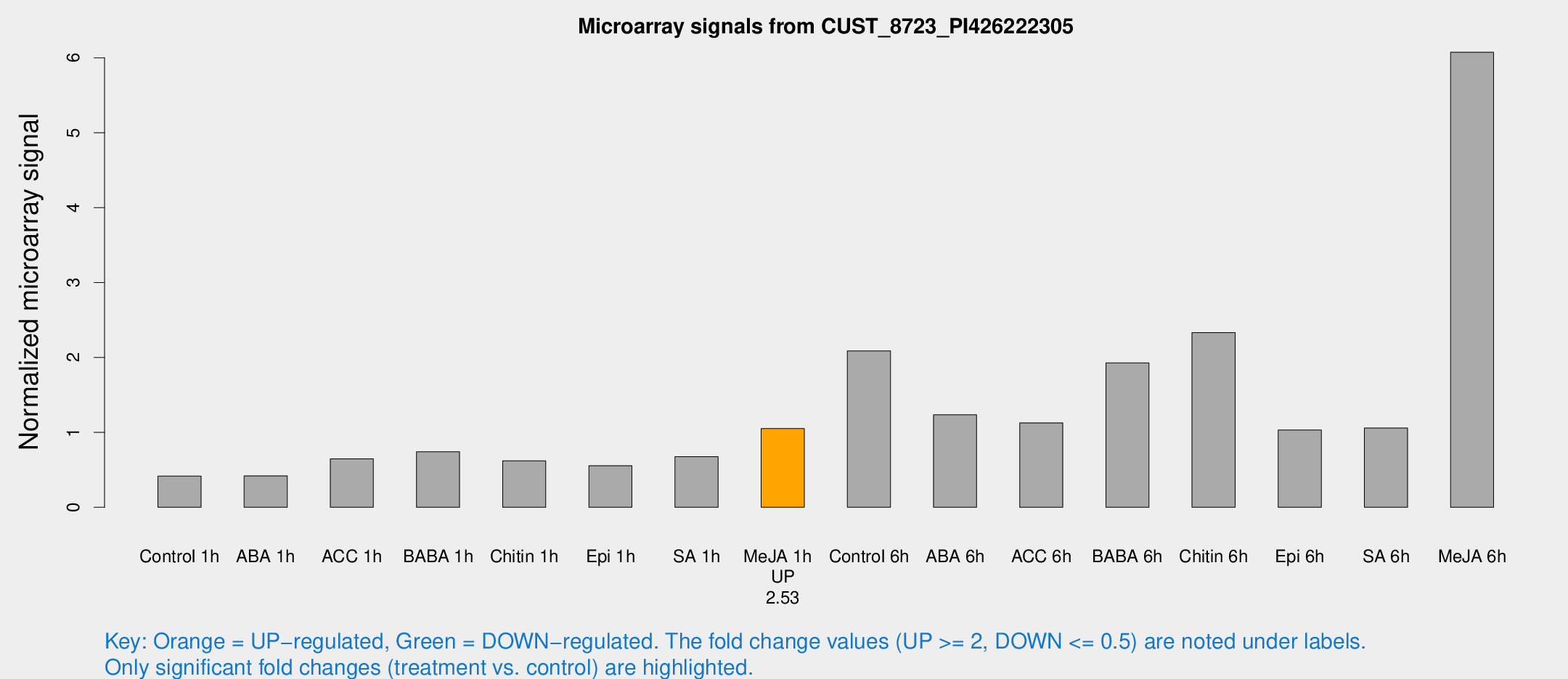
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 366.484 | 22.0702 | 0.415942 | 0.0417657 |
| ABA 1h | 362.519 | 109.218 | 0.418585 | 0.123408 |
| ACC 1h | 633.434 | 185.641 | 0.647905 | 0.148005 |
| BABA 1h | 628.922 | 36.5839 | 0.741019 | 0.0802351 |
| Chitin 1h | 491.079 | 28.6026 | 0.620933 | 0.0660782 |
| Epi 1h | 447.098 | 97.5759 | 0.554341 | 0.139997 |
| SA 1h | 651.037 | 175.9 | 0.67515 | 0.190415 |
| Me-JA 1h | 764.407 | 90.9308 | 1.05153 | 0.0608988 |
| Control 6h | 2409.21 | 1234.14 | 2.08835 | 1.06639 |
| ABA 6h | 1157.45 | 74.2621 | 1.23463 | 0.149934 |
| ACC 6h | 1227.05 | 332.889 | 1.12737 | 0.229842 |
| BABA 6h | 1921.99 | 233.646 | 1.92856 | 0.216625 |
| Chitin 6h | 2517.34 | 999.765 | 2.33294 | 0.967353 |
| Epi 6h | 1215.08 | 404.485 | 1.0317 | 0.675838 |
| SA 6h | 948.076 | 165.618 | 1.05855 | 0.105386 |
| Me-JA 6h | 5850.85 | 1614.37 | 6.07441 | 1.56284 |
Source Transcript PGSC0003DMT400071690 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G45310.1 | +1 | 1e-170 | 487 | 233/334 (70%) | Cysteine proteinases superfamily protein | chr3:16628704-16630473 REVERSE LENGTH=358 |
