Probe CUST_8397_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_8397_PI426222305 | JHI_St_60k_v1 | DMT400039929 | ACTCTCAAGAGTCCAAAGATTTGATGAAGAATTTGGCCCATTTCTTCAATGTATTAGCCT |
All Microarray Probes Designed to Gene DMG400015446
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_8397_PI426222305 | JHI_St_60k_v1 | DMT400039929 | ACTCTCAAGAGTCCAAAGATTTGATGAAGAATTTGGCCCATTTCTTCAATGTATTAGCCT |
Microarray Signals from CUST_8397_PI426222305
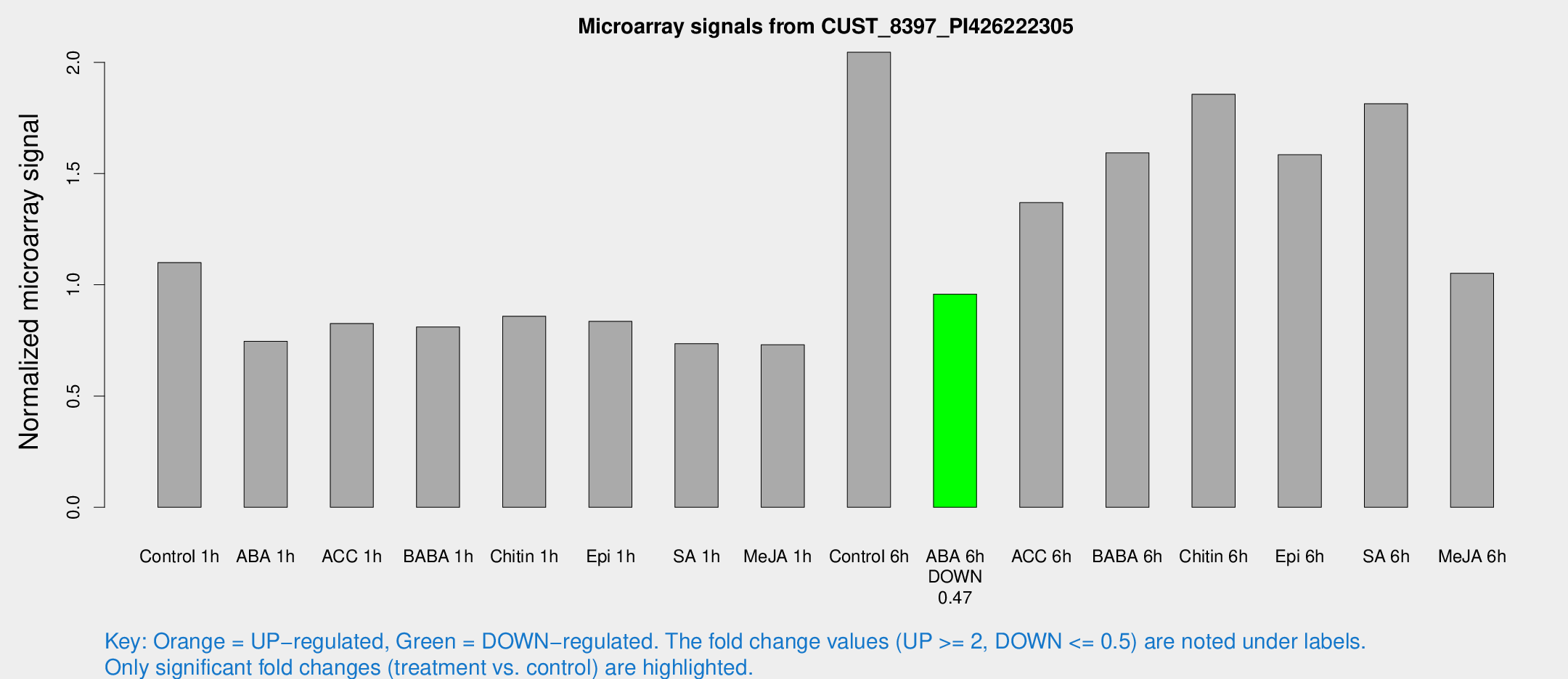
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 689.283 | 96.2256 | 1.09993 | 0.102838 |
| ABA 1h | 412.557 | 54.2224 | 0.746002 | 0.0672376 |
| ACC 1h | 533.75 | 81.7735 | 0.82619 | 0.0709611 |
| BABA 1h | 502.002 | 98.986 | 0.810504 | 0.0979004 |
| Chitin 1h | 477.944 | 44.5572 | 0.858765 | 0.0535683 |
| Epi 1h | 452.188 | 63.4673 | 0.835938 | 0.0936034 |
| SA 1h | 473.433 | 72.0958 | 0.73554 | 0.0735527 |
| Me-JA 1h | 371.973 | 49.3585 | 0.730888 | 0.0474329 |
| Control 6h | 1295.44 | 214.968 | 2.0453 | 0.206608 |
| ABA 6h | 625.795 | 36.39 | 0.958382 | 0.0556426 |
| ACC 6h | 999.647 | 179.236 | 1.36971 | 0.176069 |
| BABA 6h | 1116.22 | 163.934 | 1.59316 | 0.206992 |
| Chitin 6h | 1239.89 | 192.434 | 1.85648 | 0.221731 |
| Epi 6h | 1104.28 | 100.392 | 1.58451 | 0.156039 |
| SA 6h | 1114.26 | 121.156 | 1.81349 | 0.104913 |
| Me-JA 6h | 719.35 | 214.404 | 1.05208 | 0.302693 |
Source Transcript PGSC0003DMT400039929 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G47480.1 | +1 | 5e-95 | 290 | 147/311 (47%) | alpha/beta-Hydrolases superfamily protein | chr1:17417623-17419296 FORWARD LENGTH=314 |
