Probe CUST_8392_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_8392_PI426222305 | JHI_St_60k_v1 | DMT400030112 | CCATTCTCATTGAAATACAGGCAAGAATTTTCGTATACGTATGTTGCACGTGTATATCAA |
All Microarray Probes Designed to Gene DMG400011538
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_8392_PI426222305 | JHI_St_60k_v1 | DMT400030112 | CCATTCTCATTGAAATACAGGCAAGAATTTTCGTATACGTATGTTGCACGTGTATATCAA |
| CUST_8094_PI426222305 | JHI_St_60k_v1 | DMT400030111 | TGCCACCATTCTCATTGAAATACAGGGCAGTAGTAAGGTCAAATTGTTTTCACAAGGAGG |
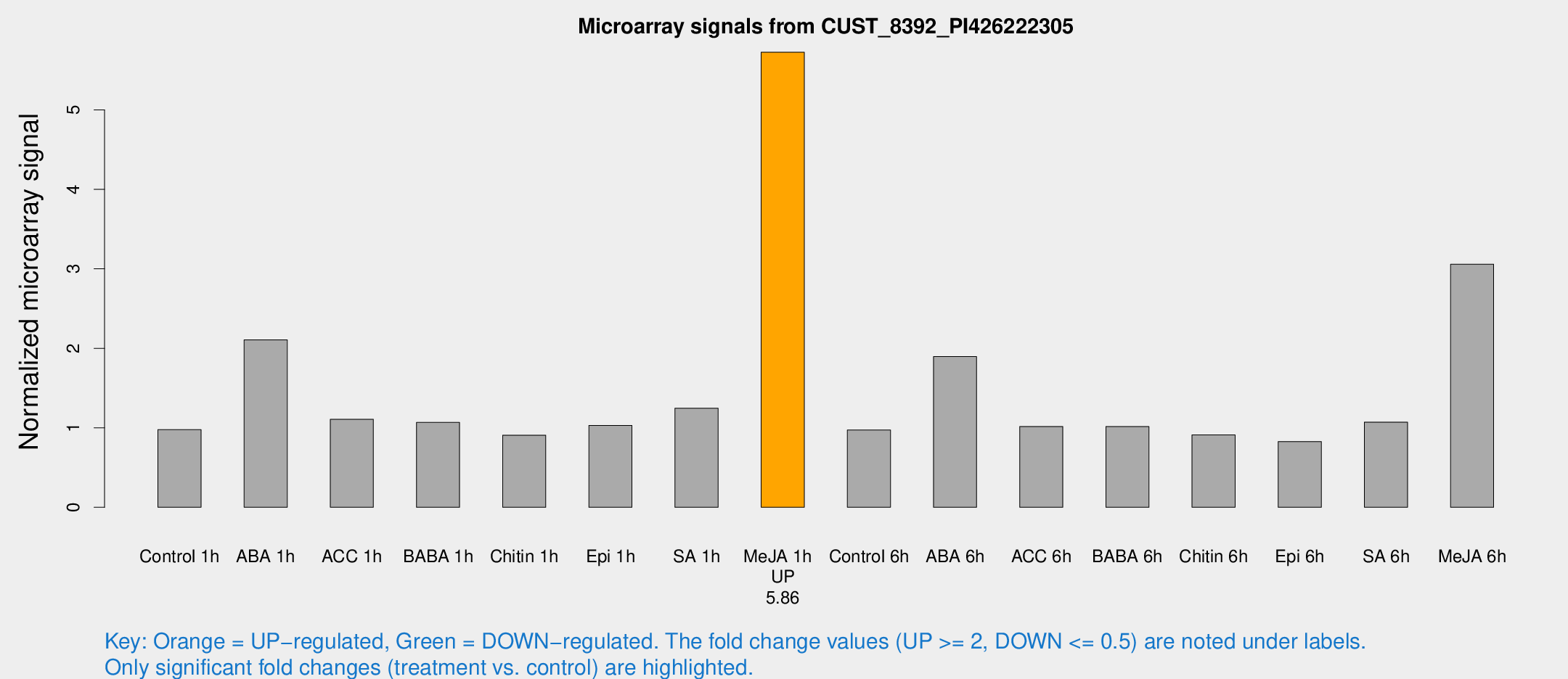
Microarray Signals from CUST_8392_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6.60926 | 2.95715 | 0.976834 | 0.474876 |
| ABA 1h | 13.593 | 4.84576 | 2.10623 | 0.801475 |
| ACC 1h | 7.98101 | 3.24248 | 1.10677 | 0.528994 |
| BABA 1h | 6.74126 | 3.14894 | 1.06793 | 0.509271 |
| Chitin 1h | 5.1617 | 3.00969 | 0.907488 | 0.510226 |
| Epi 1h | 5.84617 | 2.9696 | 1.03042 | 0.53531 |
| SA 1h | 10.6128 | 5.42389 | 1.24695 | 0.705729 |
| Me-JA 1h | 30.3953 | 3.53006 | 5.72518 | 0.666587 |
| Control 6h | 6.51054 | 3.15015 | 0.972494 | 0.496 |
| ABA 6h | 14.8181 | 5.08325 | 1.89567 | 0.670942 |
| ACC 6h | 7.66949 | 3.78966 | 1.01657 | 0.496854 |
| BABA 6h | 7.64234 | 3.55264 | 1.01606 | 0.494357 |
| Chitin 6h | 6.33451 | 3.44625 | 0.91018 | 0.499435 |
| Epi 6h | 6.119 | 3.57507 | 0.827615 | 0.479317 |
| SA 6h | 7.22472 | 3.31512 | 1.0719 | 0.535503 |
| Me-JA 6h | 20.3204 | 3.37989 | 3.05855 | 0.581304 |
Source Transcript PGSC0003DMT400030112 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G49630.1 | +1 | 7e-126 | 374 | 194/259 (75%) | amino acid permease 6 | chr5:20142681-20146441 REVERSE LENGTH=481 |
