Probe CUST_8351_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_8351_PI426222305 | JHI_St_60k_v1 | DMT400075590 | GGTATCAAATGAAGTTGTAATCACTAAATTTTGTGGGGAGAAGATAGTTTTAGCTTGGTG |
All Microarray Probes Designed to Gene DMG400029396
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_8202_PI426222305 | JHI_St_60k_v1 | DMT400075589 | GGTATCAAATGAAGTTGTAATCACTAAATTTTGTGGGGAGAAGATAGTTTTAGCTTGGTG |
| CUST_8351_PI426222305 | JHI_St_60k_v1 | DMT400075590 | GGTATCAAATGAAGTTGTAATCACTAAATTTTGTGGGGAGAAGATAGTTTTAGCTTGGTG |
Microarray Signals from CUST_8351_PI426222305
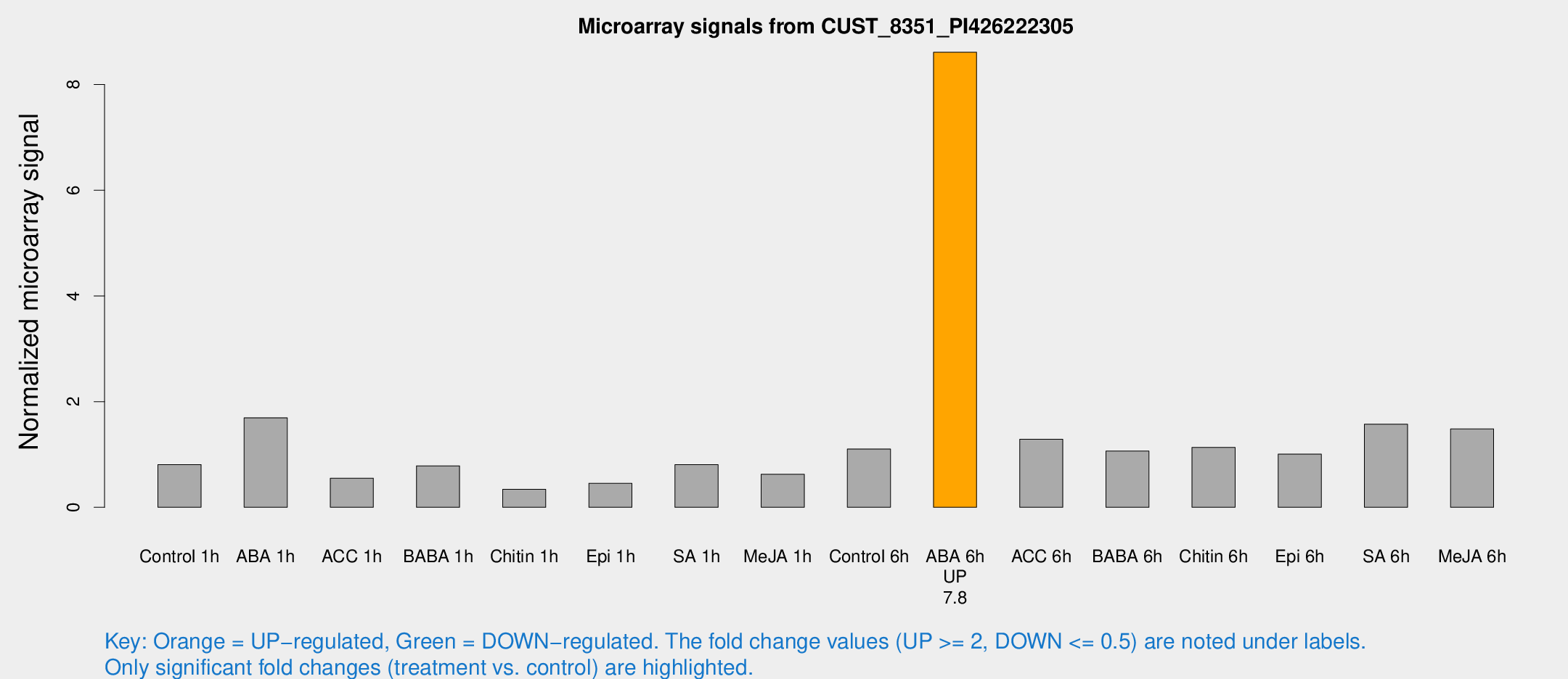
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 48.0999 | 5.02897 | 0.80916 | 0.0848955 |
| ABA 1h | 95.3145 | 26.7867 | 1.69404 | 0.466946 |
| ACC 1h | 38.8296 | 12.4126 | 0.550753 | 0.213735 |
| BABA 1h | 46.6506 | 9.65772 | 0.784557 | 0.101002 |
| Chitin 1h | 22.4013 | 10.8762 | 0.340813 | 0.144441 |
| Epi 1h | 29.4515 | 11.3492 | 0.453804 | 0.309338 |
| SA 1h | 50.6137 | 9.3911 | 0.807896 | 0.146673 |
| Me-JA 1h | 30.428 | 3.73381 | 0.624735 | 0.0774264 |
| Control 6h | 68.0766 | 15.2023 | 1.1034 | 0.212011 |
| ABA 6h | 554.928 | 92.8922 | 8.61183 | 1.06374 |
| ACC 6h | 87.4561 | 7.1183 | 1.28724 | 0.286832 |
| BABA 6h | 70.2786 | 5.32367 | 1.06456 | 0.0805043 |
| Chitin 6h | 71.3416 | 5.66553 | 1.13231 | 0.0854582 |
| Epi 6h | 77.651 | 28.0135 | 1.00545 | 0.36004 |
| SA 6h | 95.0482 | 18.291 | 1.57322 | 0.168584 |
| Me-JA 6h | 89.8277 | 16.7158 | 1.48325 | 0.151805 |
Source Transcript PGSC0003DMT400075590 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G52190.1 | +1 | 1e-162 | 488 | 269/553 (49%) | Major facilitator superfamily protein | chr1:19434671-19438673 FORWARD LENGTH=607 |
