Probe CUST_8009_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_8009_PI426222305 | JHI_St_60k_v1 | DMT400075422 | CTCATTGACTCAAAAAATGGTAGACTTCAAATTGAGCTACAATGGAGAACTGCTTCATGA |
All Microarray Probes Designed to Gene DMG400029335
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_8009_PI426222305 | JHI_St_60k_v1 | DMT400075422 | CTCATTGACTCAAAAAATGGTAGACTTCAAATTGAGCTACAATGGAGAACTGCTTCATGA |
Microarray Signals from CUST_8009_PI426222305
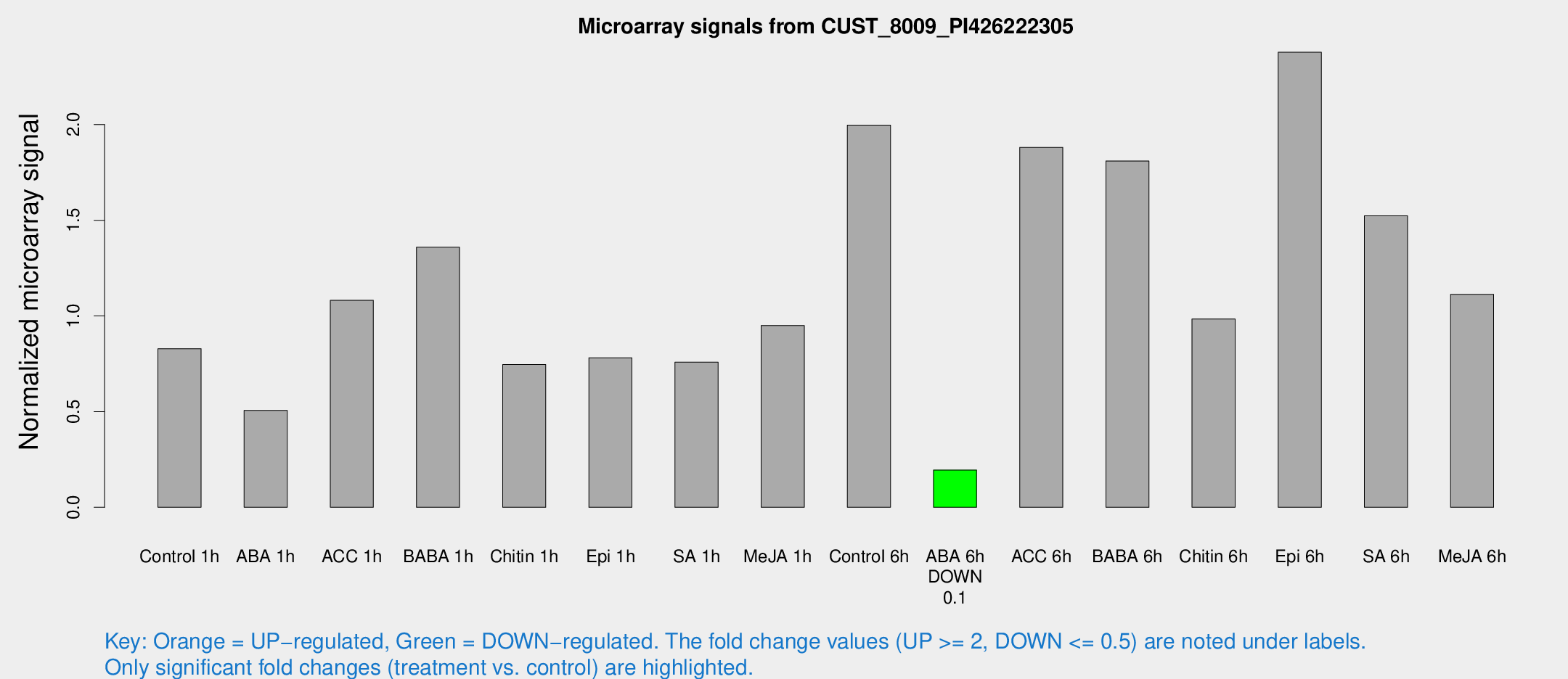
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 31.1482 | 11.7161 | 0.829107 | 0.328868 |
| ABA 1h | 14.9416 | 3.56755 | 0.506484 | 0.131102 |
| ACC 1h | 37.1693 | 6.80956 | 1.08206 | 0.29414 |
| BABA 1h | 42.6704 | 4.45016 | 1.35944 | 0.250378 |
| Chitin 1h | 26.9287 | 12.6619 | 0.745898 | 0.382399 |
| Epi 1h | 26.1369 | 11.3706 | 0.781399 | 0.389193 |
| SA 1h | 25.7303 | 3.9267 | 0.758484 | 0.131382 |
| Me-JA 1h | 28.0382 | 9.26413 | 0.950079 | 0.312584 |
| Control 6h | 76.6137 | 25.1858 | 1.99737 | 0.785587 |
| ABA 6h | 6.71026 | 3.88868 | 0.194928 | 0.112907 |
| ACC 6h | 104.885 | 55.6444 | 1.88111 | 1.86096 |
| BABA 6h | 65.9847 | 5.67569 | 1.80988 | 0.214819 |
| Chitin 6h | 36.86 | 10.4683 | 0.984405 | 0.268425 |
| Epi 6h | 96.1638 | 32.0694 | 2.37839 | 0.846709 |
| SA 6h | 50.2131 | 8.88792 | 1.52391 | 0.232191 |
| Me-JA 6h | 42.8757 | 18.3141 | 1.11302 | 0.484715 |
Source Transcript PGSC0003DMT400075422 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G20080.1 | +1 | 0.0 | 795 | 371/539 (69%) | Calcium-dependent lipid-binding (CaLB domain) family protein | chr1:6962236-6964912 FORWARD LENGTH=537 |
