Probe CUST_8005_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_8005_PI426222305 | JHI_St_60k_v1 | DMT400090218 | TAAAAAACCATGGTTGATTGGTCCTTTGTTTCACTTTTCAAAGAGAGAAGAAGCAAGCAA |
All Microarray Probes Designed to Gene DMG400039789
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_8005_PI426222305 | JHI_St_60k_v1 | DMT400090218 | TAAAAAACCATGGTTGATTGGTCCTTTGTTTCACTTTTCAAAGAGAGAAGAAGCAAGCAA |
Microarray Signals from CUST_8005_PI426222305
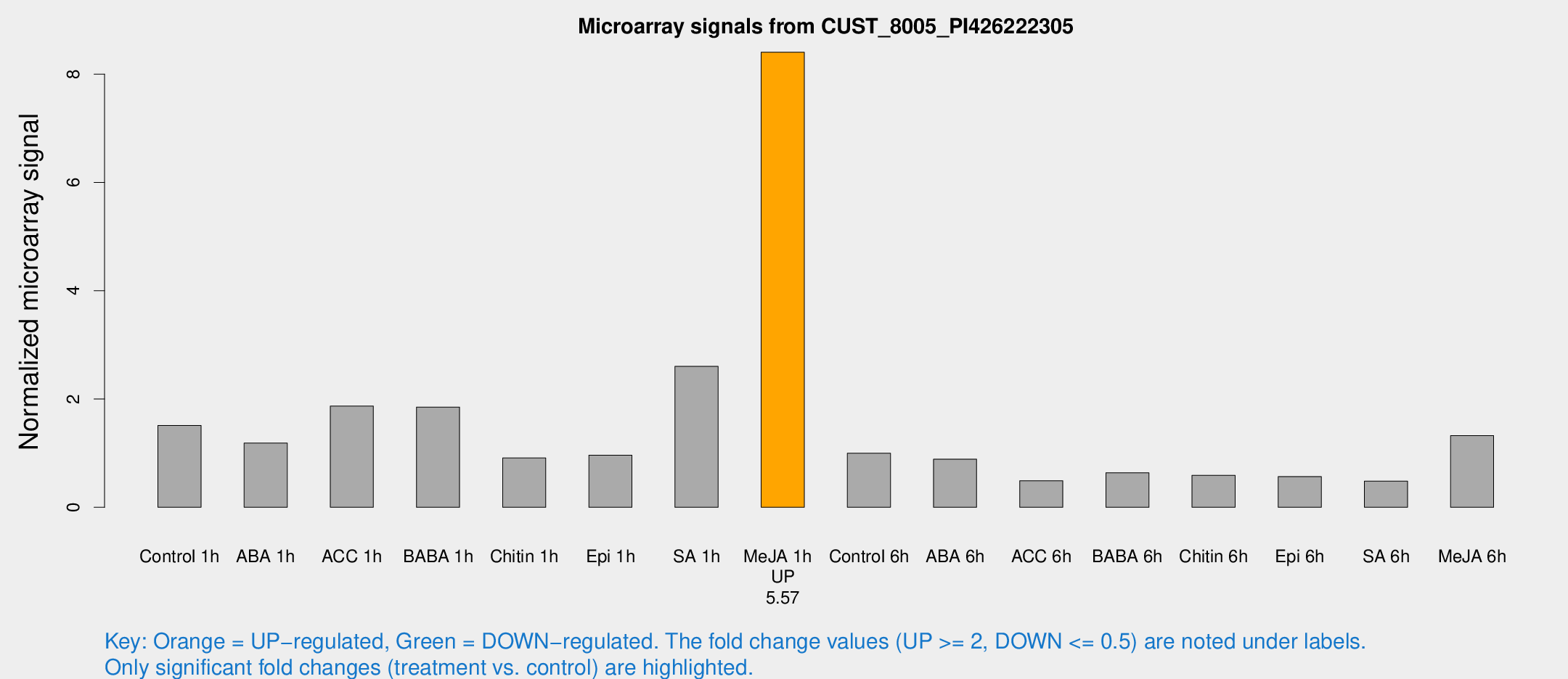
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 174.581 | 58.3265 | 1.51008 | 0.505419 |
| ABA 1h | 124.337 | 44.1723 | 1.18547 | 0.405039 |
| ACC 1h | 245.084 | 114.605 | 1.86893 | 0.870393 |
| BABA 1h | 188.024 | 27.1248 | 1.85094 | 0.11408 |
| Chitin 1h | 86.3977 | 12.2365 | 0.911928 | 0.166182 |
| Epi 1h | 91.2128 | 21.3617 | 0.96226 | 0.249347 |
| SA 1h | 283.216 | 48.3674 | 2.60268 | 0.519105 |
| Me-JA 1h | 718.582 | 93.3303 | 8.40456 | 0.487262 |
| Control 6h | 114.035 | 34.7974 | 0.998265 | 0.244777 |
| ABA 6h | 104.958 | 26.6325 | 0.889166 | 0.213965 |
| ACC 6h | 59.0054 | 8.00771 | 0.490852 | 0.0466167 |
| BABA 6h | 76.9124 | 16.5323 | 0.638304 | 0.111308 |
| Chitin 6h | 67.2211 | 11.9395 | 0.591583 | 0.138754 |
| Epi 6h | 74.7714 | 23.0755 | 0.567601 | 0.285172 |
| SA 6h | 54.8023 | 15.9448 | 0.484126 | 0.115719 |
| Me-JA 6h | 137.052 | 12.985 | 1.32483 | 0.149778 |
Source Transcript PGSC0003DMT400090218 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G34135.1 | +1 | 3e-113 | 346 | 187/471 (40%) | UDP-glucosyltransferase 73B2 | chr4:16345476-16347016 REVERSE LENGTH=483 |
