Probe CUST_7994_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_7994_PI426222305 | JHI_St_60k_v1 | DMT400075403 | CACATCATCTTGCTACTAAAGGTAATTGATGAATGGTAGAGAAGAAGAAGAGCATGAGAT |
All Microarray Probes Designed to Gene DMG400029327
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_8152_PI426222305 | JHI_St_60k_v1 | DMT400075402 | TACCAGAGGCAACTGAAGGAGTACAAAGAAAGCATCATGACCACATCATCTTGCTACTAA |
| CUST_7994_PI426222305 | JHI_St_60k_v1 | DMT400075403 | CACATCATCTTGCTACTAAAGGTAATTGATGAATGGTAGAGAAGAAGAAGAGCATGAGAT |
Microarray Signals from CUST_7994_PI426222305
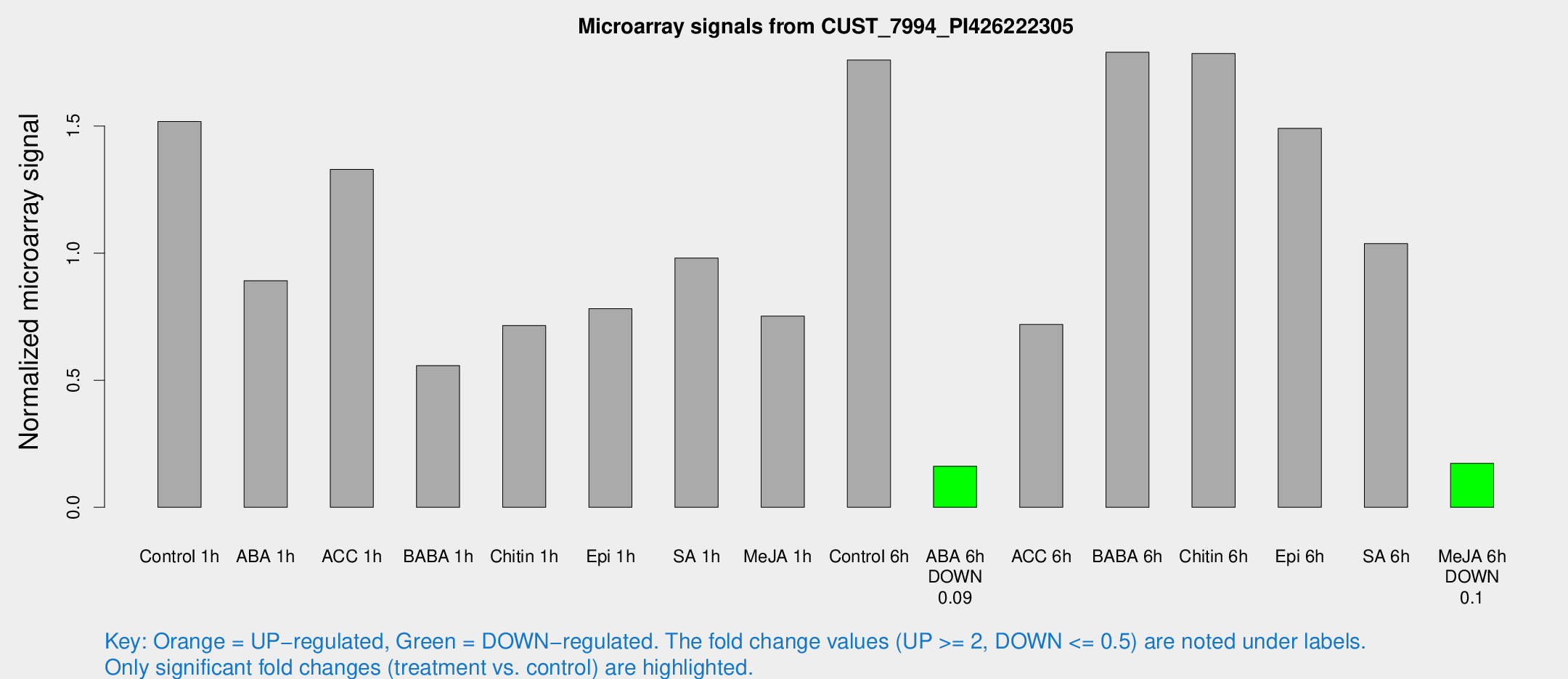
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 600.751 | 148.019 | 1.51769 | 0.272221 |
| ABA 1h | 353.214 | 148.614 | 0.891175 | 0.492069 |
| ACC 1h | 522.609 | 79.5189 | 1.32931 | 0.310459 |
| BABA 1h | 217.352 | 59.0103 | 0.557207 | 0.111056 |
| Chitin 1h | 251.827 | 55.0974 | 0.715245 | 0.152974 |
| Epi 1h | 256.305 | 28.1661 | 0.781545 | 0.0965898 |
| SA 1h | 378.739 | 22.3741 | 0.980857 | 0.0572345 |
| Me-JA 1h | 239.729 | 50.8834 | 0.752244 | 0.164596 |
| Control 6h | 713.96 | 190.553 | 1.76011 | 0.383607 |
| ABA 6h | 65.2931 | 7.69933 | 0.161771 | 0.025802 |
| ACC 6h | 386.121 | 161.118 | 0.719479 | 0.315069 |
| BABA 6h | 835.985 | 249.558 | 1.79049 | 0.724865 |
| Chitin 6h | 740.196 | 152.001 | 1.78547 | 0.29673 |
| Epi 6h | 640.417 | 78.5983 | 1.49114 | 0.302479 |
| SA 6h | 424.112 | 115.228 | 1.03749 | 0.231068 |
| Me-JA 6h | 80.8665 | 35.8317 | 0.172554 | 0.0810014 |
Source Transcript PGSC0003DMT400075403 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G76110.1 | +2 | 1e-90 | 281 | 161/300 (54%) | HMG (high mobility group) box protein with ARID/BRIGHT DNA-binding domain | chr1:28555287-28557465 REVERSE LENGTH=338 |
