Probe CUST_6946_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_6946_PI426222305 | JHI_St_60k_v1 | DMT400031252 | TGCAACAAATGATGGACACAATGGACAGAGTCATGGAGGACCCTCTTGCTTTTAATGGAG |
All Microarray Probes Designed to Gene DMG400011977
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_6946_PI426222305 | JHI_St_60k_v1 | DMT400031252 | TGCAACAAATGATGGACACAATGGACAGAGTCATGGAGGACCCTCTTGCTTTTAATGGAG |
| CUST_6999_PI426222305 | JHI_St_60k_v1 | DMT400031253 | GAGGTTAAAGATGGTGTCTTGTATATTACTATACCTAAGGCTAGTAGTAATCCCAAAGTT |
Microarray Signals from CUST_6946_PI426222305
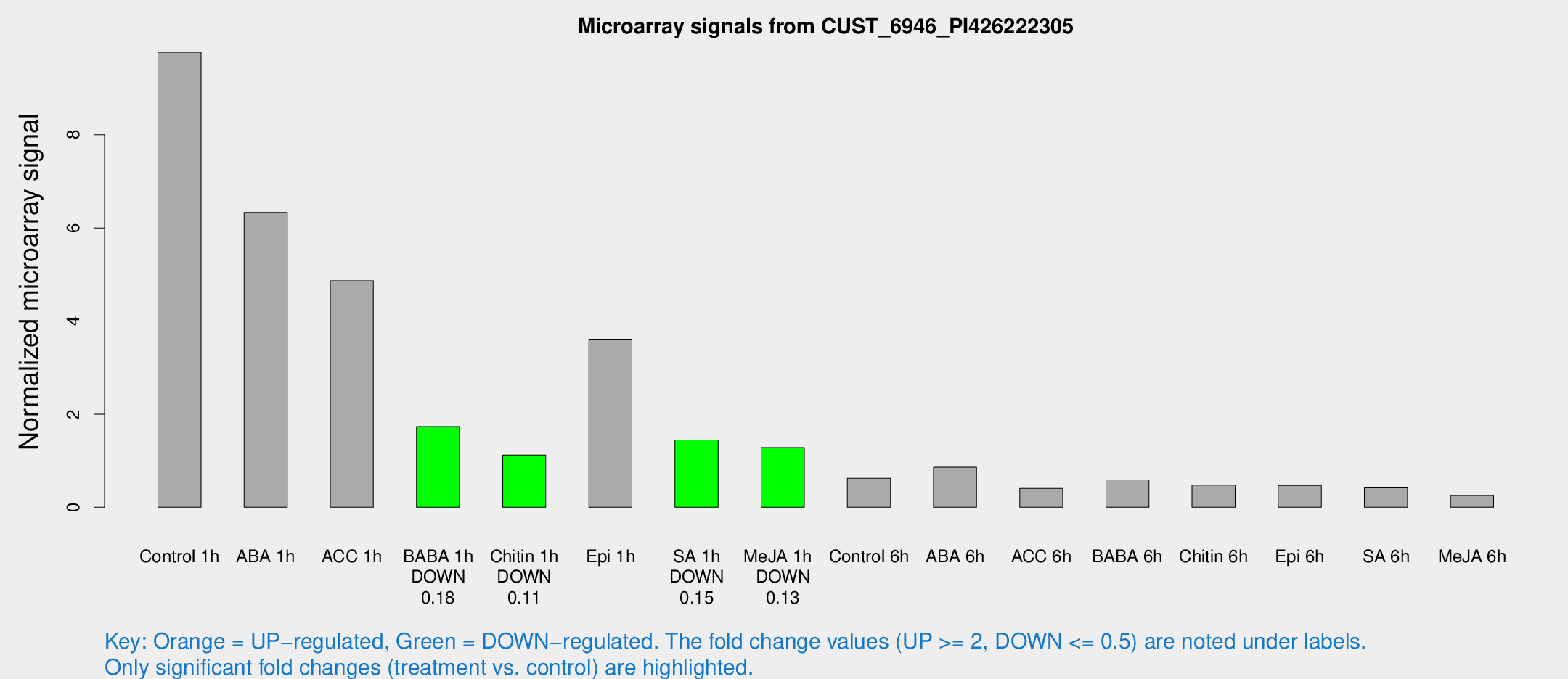
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 488.543 | 76.782 | 9.77272 | 1.40422 |
| ABA 1h | 418.076 | 263.939 | 6.33251 | 6.29859 |
| ACC 1h | 344.012 | 177.924 | 4.86686 | 3.52524 |
| BABA 1h | 90.1975 | 30.2126 | 1.73314 | 0.473325 |
| Chitin 1h | 50.3682 | 7.29333 | 1.12307 | 0.101591 |
| Epi 1h | 167.177 | 46.125 | 3.59722 | 1.39162 |
| SA 1h | 79.0778 | 23.7652 | 1.44631 | 0.420439 |
| Me-JA 1h | 51.7097 | 5.74976 | 1.28177 | 0.216954 |
| Control 6h | 31.4955 | 5.70121 | 0.622932 | 0.0867475 |
| ABA 6h | 44.8389 | 4.49047 | 0.860007 | 0.112055 |
| ACC 6h | 22.8598 | 4.48133 | 0.404771 | 0.0768878 |
| BABA 6h | 32.2674 | 4.33752 | 0.587471 | 0.0789668 |
| Chitin 6h | 24.7075 | 4.11319 | 0.475479 | 0.0793453 |
| Epi 6h | 26.2978 | 4.34503 | 0.470127 | 0.0887848 |
| SA 6h | 21.773 | 5.26158 | 0.418311 | 0.169308 |
| Me-JA 6h | 12.742 | 3.63569 | 0.253411 | 0.0784578 |
Source Transcript PGSC0003DMT400031252 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G27670.1 | +2 | 7e-43 | 148 | 86/230 (37%) | heat shock protein 21 | chr4:13819048-13819895 REVERSE LENGTH=227 |
