Probe CUST_6571_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_6571_PI426222305 | JHI_St_60k_v1 | DMT400014323 | AACAAGTGGATAGAAATGATGATAATGTTGTAACCGGGCCAAAAGGGCACCATCATGTTT |
All Microarray Probes Designed to Gene DMG400005620
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_6571_PI426222305 | JHI_St_60k_v1 | DMT400014323 | AACAAGTGGATAGAAATGATGATAATGTTGTAACCGGGCCAAAAGGGCACCATCATGTTT |
Microarray Signals from CUST_6571_PI426222305
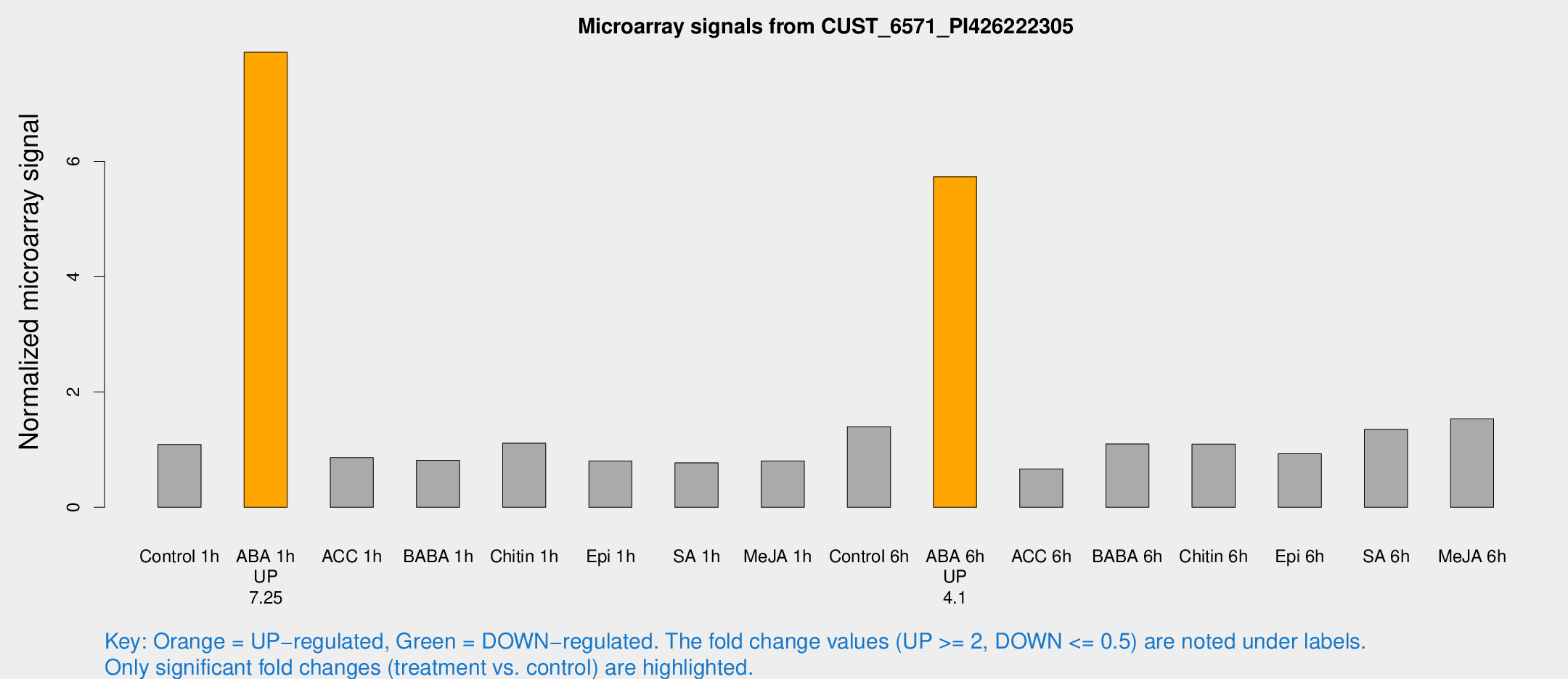
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 37.5103 | 9.17577 | 1.08919 | 0.339249 |
| ABA 1h | 229.842 | 22.0453 | 7.8924 | 0.471061 |
| ACC 1h | 31.2076 | 9.07959 | 0.861884 | 0.331636 |
| BABA 1h | 25.8289 | 3.94988 | 0.81666 | 0.12658 |
| Chitin 1h | 32.9482 | 3.95939 | 1.11221 | 0.1367 |
| Epi 1h | 22.688 | 3.63305 | 0.802601 | 0.129837 |
| SA 1h | 27.493 | 6.78357 | 0.771792 | 0.180517 |
| Me-JA 1h | 22.9196 | 6.27647 | 0.802998 | 0.30264 |
| Control 6h | 49.1055 | 13.8883 | 1.39831 | 0.360315 |
| ABA 6h | 200.254 | 18.3015 | 5.73242 | 0.347808 |
| ACC 6h | 25.0061 | 4.67467 | 0.665162 | 0.129521 |
| BABA 6h | 40.3608 | 4.57826 | 1.09851 | 0.134922 |
| Chitin 6h | 38.6379 | 5.01571 | 1.09428 | 0.164835 |
| Epi 6h | 35.8955 | 8.41183 | 0.927558 | 0.145176 |
| SA 6h | 43.5487 | 4.54772 | 1.35036 | 0.184183 |
| Me-JA 6h | 56.3298 | 18.3972 | 1.53499 | 0.451388 |
Source Transcript PGSC0003DMT400014323 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G18570.1 | +3 | 4e-42 | 145 | 95/167 (57%) | Oleosin family protein | chr3:6396072-6396572 REVERSE LENGTH=166 |
